Pooled analysis of 3,741 faecal metagenomes from 18 cohorts for the identification of reproducible microbial biomarkers at different stages of colorectal cancer
By Prof Harry Sokol
Gastroenterology and Nutrition Department,
Saint-Antoine Hospital, Paris, France
Commentary on the article by Piccinno et al. (Nature Medicine 2025 1)
Lay public section
Find here your dedicated section
Sources
This article is based on scientific information
Sections
About this article
Associations between the gut microbiome and colorectal cancer (CRC) have been uncovered, but larger and more diverse studies are needed to assess their potential clinical use. The authors of this article used 12 metagenomic datasets of patients with CRC (n = 930), adenomas (n = 210) and healthy controls (n = 976; total n = 2,116) and added six new cohorts (n = 1,625) providing granular information on cancer stage and the anatomic location of the tumours. They improved CRC prediction accuracy based solely on gut metagenomics (average area under the curve = 0.85) and highlighted the contribution of 19 new species and distinct Fusobacterium nucleatum clades. Specific gut species distinguish left-sided versus right-sided CRC (area under the curve = 0.66) with an enrichment of oral-typical microbes. The authors identified strain-specific CRC signatures with the commensal species of Ruminococcus bicirculans and Faecalibacterium prausnitzii, showing subclades associated with advanced CRC. The analysis confirmed that the microbiome can be a clinical target for CRC screening and characterised it as a biomarker for CRC progression.
What do we already know about this subject?
CRC is the third most frequent and the second most lethal tumour type worldwide 2. CRC originates in the epithelial layer of the proximal colon (right colon) or distal colon plus rectum (left colon). Progression from benign precancerous lesion (adenoma) to a malignant tumour (carcinoma) may take several years and is characterised by an accumulation of tumour-cell mutations, alteration in the gut mucosal barrier and intestinal inflammation.
The gut microbiome is proposed as one of the important hallmarks of cancer. Certain microbes have been put forward as major contributors to carcinogenesis, particularly Escherichia coli pks+ and Fusobacterium nucleatum 3. Several studies have observed distinct microbiome signatures in CRC patients when compared with patients with adenomas or healthy controls 4. A few metagenomic studies also investigated
microbiome changes along the adenoma-carcinoma sequence and based on the primary neoplasia location, and have suggested links between CRC and oral species. Further evidence points toward the enrichment of oral-typical microbes and of oral biofilm-forming species
in the gut metagenomes of patients with proximal CRC. However, no metagenomic studies have gone beyond characterising already well-known strain-specific factors that influence CRC risk, and no untargeted searches for subspecies and strain-level genomic associations with CRC phenotypes are available.
What are the main insights from this study?
Using 3,741 samples from 18 cohorts and applying new strain-level computational methodologies, the authors investigated the links between faecal microbiota and CRC. They improved CRC prediction accuracy based solely on gut metagenomics, with an average area under the curve (AUC) = 0.85. The five SGBs (Species- level Genome Bins) assigned to the species F. nucleatum were more abundant
in CRC than in the controls: F. nucleatum subsp. animalis, vincentii, nucleatum, polymorphum. This was in addition to other well-characterised CRC-associated microbes such as Parvimonas micra and Bacteroides fragilis. The authors also identified 19 additional uncharacterised SGBs with neither cultivated strains nor taxonomically defined species, which highlighted a more complex CRC-associated
microbial signature than previously thought.
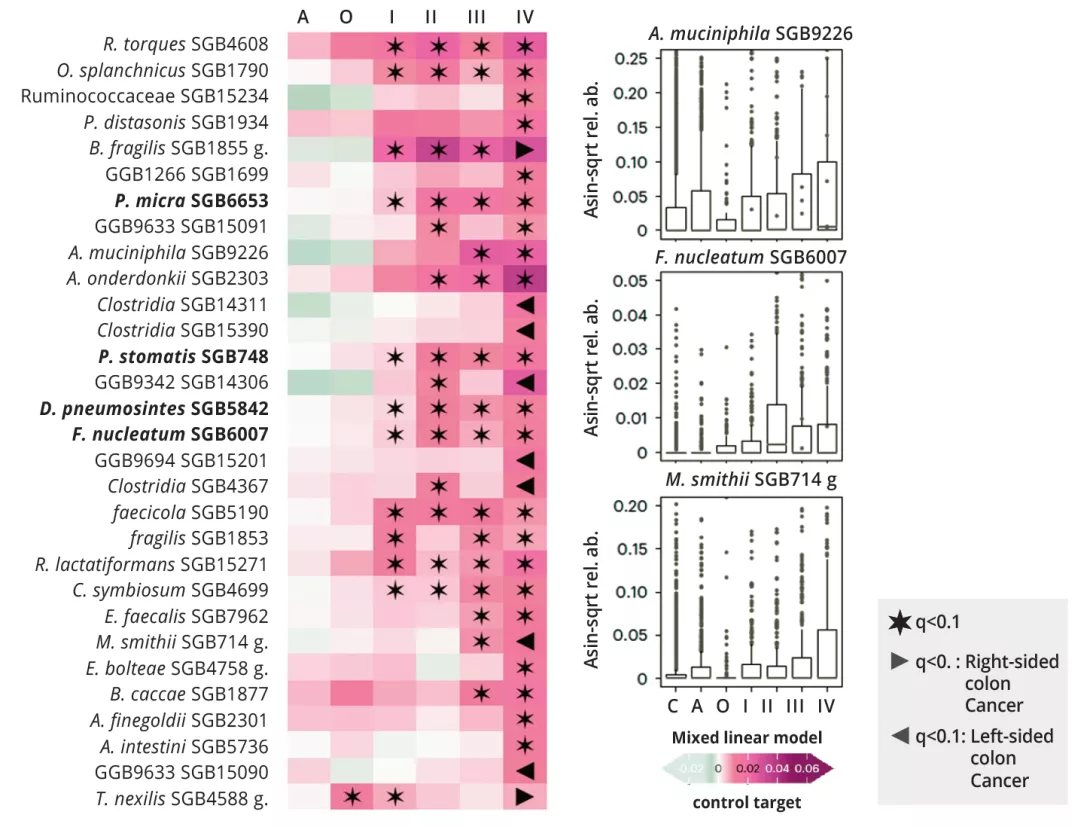
Although interstage microbiome variations during CRC progression are not as strong as those observed between CRC and controls, the authors found several biomarkers for advanced and metastatic CRC, as well as several microbial species consistently and monotonically increasing (or decreasing) from control to cancer or advanced disease. In particular, latestage CRC was enriched in oral-derived species, such as P. micra, already involved in the stimulation of tissue invasion pathways and Hungatella hathewayi, which was shown to promote intestinal cell proliferation in in-vitro experiments (figure 1). Compared with the other stages, metastatic CRC had a higher abundance of Methanobrevibacter smithii, supporting previous findings linking methane producers with stage IV CRC. Stool samples from patients with CRC originating in the right or transverse colon were also enriched in oral species.
These findings strengthen the notion that the number and cumulative abundance of orally derived species are significantly higher in CRC samples than controls and adenomas, but also show that later stages of CRC were particularly enriched in oral species. However, many non-oral bacteria were also associated with CRC, including species previously associated with high cardiometabolic risk. Interestingly, adenoma and later cancer stages were enriched in species linked with poor cardiometabolic health and immune-mediated diseases.
What are the consequences in practice?
This is the largest and most accurate study ever carried out on faecal microbiota associated with CRC. A signature based solely on faecal microbiota offers a relatively good predictive value. In addition, differences between early and late stages have been identified. However, it should be noted that the study did not identify sufficiently accurate markers for the diagnosis of pre-cancerous lesions (adenoma). This work paves the way for microbiota-based tests for diagnosing CRC, but their use in clinical practice still requires validation and, above all, the detection of pre-cancerous lesions needs to be improved.
Figure 1 : Microbial signature according to CRC stage and location of primary tumour.
- Using 3,741 samples from 18 cohorts, the authors investigated the links between faecal microbiota and CRC
- A signature based solely on faecal microbiota provides a relatively good predictive value, with an average area under the curve (AUC) = 0.85
- However, the study did not identify sufficiently accurate markers for the diagnosis of pre cancerous lesions
- Alongside differences between control and CRC subjects, alterations in the microbiota were observed according to the stage and topography of the primary lesion, particularly in terms of the abundance of oral bacteria
CONCLUSION
A signature based solely on faecal microbiota offers prediction with an average area under the curve (AUC) = 0.85. However, the study did not identify sufficiently accurate markers for the diagnosis of pre-cancerous lesions (adenomas). In addition to differences between control and CRC subjects, alterations in the microbiota were observed according to the stage and topography of the primary lesion, particularly in terms of oral bacteria abundance.

