Targeting the gut microbiota to combat allergies
Overview
By Pr. Roberto Berni Canani, MD, PhD
Department of Translational Medical Science; European Laboratory for the Investigation of Food Induced Diseases, and CEINGE Advanced Biotechnologies; Task Force on Investigation on Microbiome, University of Naples “Federico II”, Naples, Italy
Lay public section
Find here your dedicated section
Sources
This article is based on scientific information
Sections
About this article
Author
Dysbiotic gut microbiota plays an important role in the development of allergic diseases, in particular food allergies. The gut microbiota drives the maturation and function of the immune system, and genetic, environmental, and dietary factors may alter the commensal microbiota, leading to a dysregulation of immune function. Several factors responsible for dysbiosis have been associated with the occurrence of allergies, such as Caesarean delivery, the lack of breast milk, the use of drugs (mainly antibiotics and gastric acidity inhibitors), the use of antiseptic agents, and low-fibre/high-fat diets. No specific bacterial taxa have been consistently associated with allergies, but evidence suggests that gut dysbiosis occurs even before allergies present. Shortchain fatty acids (SCFAs) are crucial gut microbiota-derived metabolites involved in cross-talk with the immune system. Targeting the composition and function of the gut microbiota represents a promising strategy against allergic diseases, in particular, against childhood food allergies.
The prevalence, persistence, and severity of allergic diseases and, in particular, food allergies (FA), has increased substantially in recent decades in the industrialised world under the pressure of gene-environment interactions leading to immune system dysfunction, mediated, at least in part, by epigenetic mechanisms [1, 2]. This changing scenario has led to an increase in hospital admissions, medical visits, treatments, and a greater burden of care on families. All these factors have a significant impact on social costs and quality of life, and place a great psychological burden on patients and families.
Food allergies are characterised by an abnormal immune response towards dietary antigenic peptides that are normally tolerated. The cause of FAs is still largely undefined. Based on current knowledge, genetic susceptibility alone cannot account for the changing pattern of FAs and there has been a renewed interest in the role of the environment in the sensitisation to food. Evidence suggests a key pathogenetic role of gut microbiota (GM) alterations (dysbiosis) in allergy development. A healthy GM has a crucial impact on the development of the gastro-intestinal tract and immune system. A dysbiotic GM is associated with various diseases, including allergies [3].
The importance of microbial exposure for immune tolerance
The way in which dietary antigens are normally rendered non-immunogenic through immune tolerance has not yet been fully defined. Evidence suggests a pivotal role for regulatory T cells (Tregs) expressing the transcription factor Foxp3 (Foxp3+Tregs) and the complex interaction between the GM and immune and non-immune cells. The presence of both diet-and microbe-induced populations of Treg cells is required for full tolerance to food antigens [4].
During vaginal delivery, infants receive their first bacterial inoculum from the maternal vaginal tract, skin tissue, and often from faecal matter, exposing the immature immune system to a significant bacterial load [3]. The maturation of a healthy GM in early life allows for a change in the Th1/Th2 balance, favouring a Th1 cell response, while dysbiosis alters host-microbiota homeostasis, producing a shift in the Th1/Th2 cytokine balance towards a Th2 response [5].
Tregs are depleted in germ-free mice and in mice receiving an amino acid-based diet [4, 6]. Secretory IgA (sIgA) and innate immunity peptides exert a pivotal role in regulating GM composition. Deficiency in both innate and adaptive immunity (especially low levels of IgA) has been observed in children with multiple FAs [7]. A healthy GM promoting sIgA production facilitates the survival of protective bacterial strains within the gut lumen [8].
The gut microbiota and allergies
The expression of an allergic phenotype is dependent on the interaction between two major factors: genetic predisposition and gene-environment interactions. An increasing number of studies suggests a correlation between factors that alter the GM in the first years of life and the development of allergies later in life. There is increasing evidence that early-life GM dysbiosis represents a critical factor underlying the development of allergies.
The main factors responsible for dysbiosis are: birth by Caesarean section, lack of breast milk, drug use (mainly antibiotics and gastric acidity inhibitors), antiseptic agent use, timing of the introduction of solid foods, and junk food-based and/or low-fibre/high-fat diets [3, 9]. The maternal use of antibiotics before and during pregnancy, as well as antibiotic treatments during the first months of life, are also reported to be associated with an increased risk of cow’s milk allergies (CMA) in children [10]. Data that may be used to characterise the microbiota of patients with FAs are still preliminary. We recently described GM dysbiosis in children affected by IgE-mediated CMA; CMA infants had significantly reduced levels of Bifidobacteriaceae, Streptococcaceae, Enterobacteriaceae, and Enterococcaceae, and presented significantly higher levels of selected strains from Ruminococcaceae and Lachnospiraceae families. The GM of subjects with CMA comprised 73% Bacteroidetes and Firmicutes taxa, which are also known to dominate the adult gut [11].
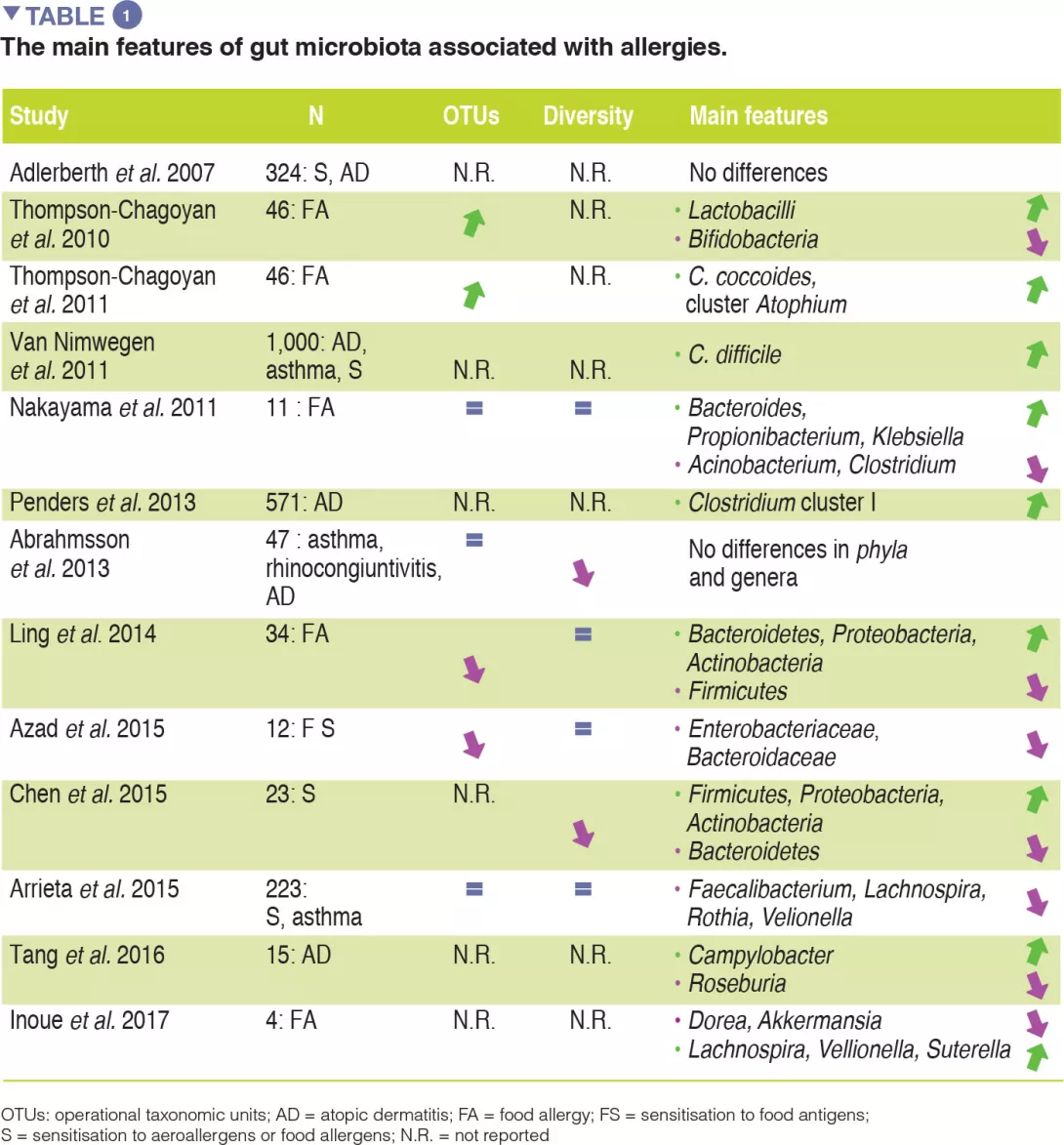
Although compelling evidence for an association between GM dysbiosis and FAs is emerging, heterogeneity in study design (involving sampling time points, the methods used to characterise microbiota, and allergic phenotypes studied) has made it difficult to establish a causal relationship between specific bacterial taxa and the development of allergies. No specific bacterial taxa have been consistently associated with FA and a broad range of microbes isolated from the human gut may be involved in tolerogenic mechanisms. The main evidence for allergy-associated GM is summarised in Table 1.
The gut microbiota plays a central role in the pathogenesis of allergies
Evidence suggests that the gut microbiota-epigenetics axis, modulated by a number of environmental and dietary factors, plays an important role in the occurrence of allergies. This axis regulates a number of non-immune and immune tolerogenic mechanisms, and could be the ideal target for innovative, preventive, and therapeutic strategies against allergies.
SCFAs are major GM metabolites involved in cross-talk with human cells. Among the SCFAs, butyrate exerts a pivotal role in the induction of immune tolerance, and butyrate deficiency has been observed in allergic patients [4]. The different types of dysbiosis may be hypothesised to lead to similar effects in terms of SCFAs, and/ or the production of other microbiota-derived metabolites may facilitate the occurrence of allergies. Clostridia species, belonging to Cluster IV and XIVa, are the prominent source of SCFAs in the colon. Bacteria-produced SCFAs have been implicated in the regulation of both the proportion and functional capability of Tregs, which, in some studies, have been specifically attributed to butyrate production by spore-forming Clostridiales. An enrichment of taxa from the Clostridia class and Firmicutes phylum has been observed in human subjects with resolution of CMA [9]. Data from our laboratory showed that oral butyrate treatment leads to a dramatic inhibition of acute allergic skin response, anaphylactic symptom scores, reduced body temperature, increased intestinal permeability, and anti-βLG lactoglobulin (BLG) IgE, IL-4, and IL-10 production in a murine model of CMA, suggesting a protective role of butyrate against FAs [9]. Butyrate exhibits multiple mechanisms of action, however, many of these involve epigenetic regulation of gene expression through the inhibition of histone deacetylase (HDAC). The inhibition of HDAC9 and 6 increases FoxP3 gene expression, as well as the production and suppressive function of Tregs [12]. We evaluated the direct effects of butyrate on peripheral blood mononuclear cells (PBMCs) from children affected by challenge-proven IgE-mediated CMA. PBMCs were stimulated with BLG in the presence or absence of butyrate. Preliminary results show that butyrate stimulates IL-10 and IFN-γ production and decreases the rate of DNA methylation of these two cytokines. The same effective butyrate dose induces demethylation of the FoxP3 promoter region and down-regulation of HDAC6/HDAC9 expression [2].
Modulation of the gut microbiota associated with allergies
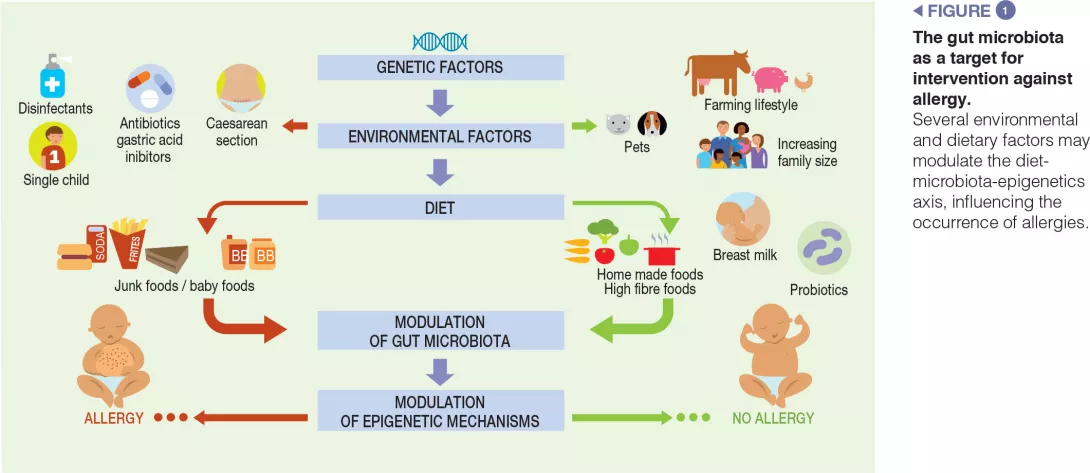
Children exposed to farm environments have a lower risk of allergy development. Although it has not been conclusively proven, one of the plausible explanations for the protective effect associated with early- life farm exposure is the role of the GM, because individuals exposed to a farm environment possess a different microbial composition relative to those with other lifestyles [3]. Other epidemiological factors which protect against FAs include older siblings and exposure to pets in early life. Pet ownership is associated with high microbial diversity in the home environment. A recent study examining the influence of dietary patterns on the development of FAs at the age of two suggests that dietary habits may influence the development of FAs by changing the composition of the gut microbiota. In particular, an infant diet consisting of high levels of fruits, vegetables, and home-prepared foods was associated with less FAs [9] (Figure 1).
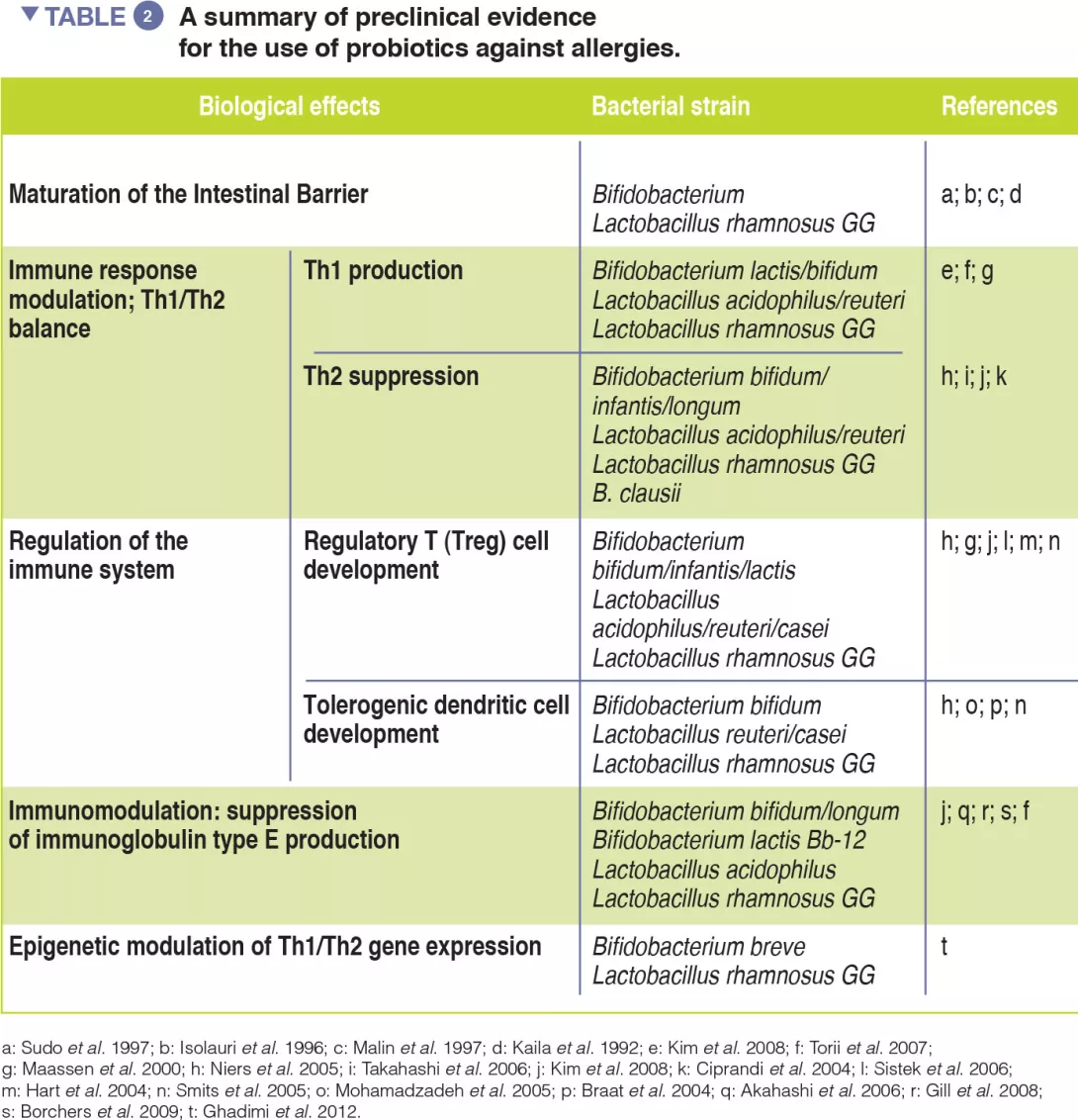
Probiotics, defined as ingested microbes that provide health benefits to the host, may be beneficial by modulating the GM [13]. Evidence of probiotic use against respiratory allergies remains preliminary (Table 2). However, meta-analyses have revealed that the use of selected probiotics, from gestation through to the first six months of life, could reduce the incidence of atopic eczema in children with a family history of allergic disease [14].
How to modulate the gut microbiota?
Emerging evidence supports the use of selected dietary strategies and probiotic strains for the prevention and treatment of allergies (Table 2).
We have previously demonstrated that addition of the probiotic, L. rhamnosus, to a hypoallergenic formula accelerates the acquisition of immune tolerance and protects against the occurrence of other atopic manifestations in children with CMA [15-17]. When we compared the faecal microbiota among infants receiving this tolerance-inducing probiotic therapy, we found significant positive correlations between the abundance of genera with a potential to produce butyrate and the concentration of faecal butyrate [11]. Strain-level demarcations for butyrate-producing genera (including Roseburia, Coprococcus, and Blautia), identified in infants who acquired tolerance to cows’ milk, suggest that L. rhamnosus treatment contributes to the acquisition of tolerance by altering the strain-level community structure of taxa with a potential to produce butyrate [11]. Accordingly, oral immunotherapy supplemented with another L. rhamnosus strain has been shown to be effective in inducing peanut non-responsiveness in 82% of allergic children [18].
Conclusion
The trillions of bacteria that populate our gut critically regulate key physiological functions against allergies. Environmentally induced changes in GM composition and function (decreased butyrate production, for example) create dysbiosis that is linked to an increased risk of allergy. Understanding how gut bacterial communities interact with the immune system is opening the way to novel preventive treatment strategies for allergies.