Crosstalk between the gut microbiota and the host's immune response to combat infections
Overview
By Dr. Dorota Czerucka
Medical Biology, Ecosystems and Immunity Team, Monaco Scientific Centre, Monaco
Lay public section
Find here your dedicated section
Sources
This article is based on scientific information
Sections
About this article
Author
The fact that living beings have evolved over millions of years in complex environments occupied by microbial ecosystems has shaped symbiotic relationships regulated by the immune system. The new sequencing techniques have revolutionised our knowledge and have shown that each individual hosts a microbiota which is unique to him, as is its role in the physiology of the host and in numerous diseases such as infections. The interaction between the gut microbiota and the immune system starts during foetal life. Their mutual and constant exchanges shape both the immunity of the host and also the gut microbiota resulting in protection from infection and numerous diseases. Indeed, the specific organisation of the microbiota - separated from the host by a single layer of cells - constitutes a particular challenge for the immune system, the role of which is to recognise “non-self” as a potential sign of infection and thus trigger the immune system cascades. For this reason, the continuous exchanges with the microbiota have a significant impact on the immune system of the host. The immune response, which must be tolerant towards the microbiota, also has an impact on the composition and function of this microbiota.
GUT MICROBIOTA AND THE INTESTINAL BARRIER
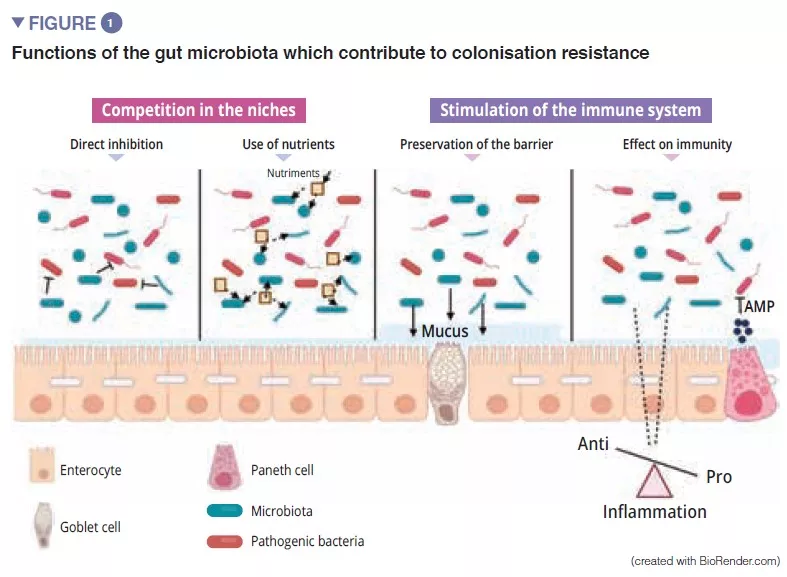
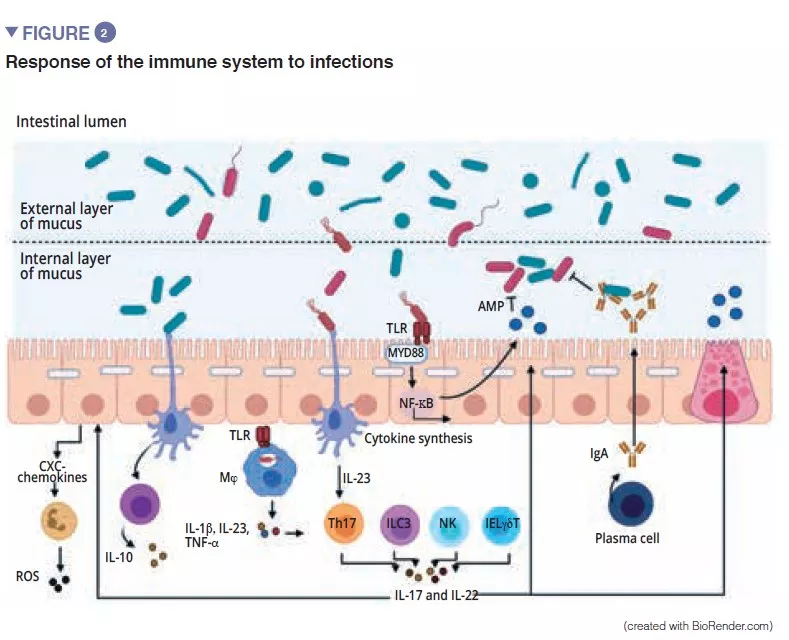
The gut microbiota is an initial barrier protecting the intestinal mucosa from pathogens. This complex ecosystem inhabits the gastrointestinal tract where it remains stable and limits access to the intestinal niches and to the nutrients required for the multiplication of exogenous bacteria by the phenomenon called “colonisation resistance” [1] (Figure 1). The enterocytes, which provide a physical barrier between the intestinal lumen and the host, absorb water and nutrients and secrete antimicrobial peptides, AMPs (RegIIIg, b-defensins and cathelicidin) [2]. By the recognition of microbe-associated molecular patterns, (MAMPs) by specific receptors (including the Toll-Like-Receptors, TLR), these cells will be able to transduce the signal to cytokines and chemokines thus signalling infection and recruiting immune cells (Figure 2). Paneth cells also participate in colonisation resistance by secreting AMPs (lysosyme, a-defensins, RegIIIg) [2]. The goblet cells – mucus-secreting – and the M cells have gatekeeping action, transporting antigens, intact and captured at random in the intestinal lumen arising from commensal bacteria or pathogens or dietary antigens. These will then be prepared by the dendritic cells (DC) and presented to the adaptive immune system. This function is vital to intestinal tolerance and the induction of mucosal immune responses [2]: there therefore is a constant balance between pro- and anti- inflammatory responses (Figure 2). In particular, this was demonstrated in mice models of induced colitis and in TLR receptor-deficient mice: the absence of microbiota or recognition of this reduces the proliferation of intestinal epithelial cells or barrier repair [2]. Lastly, the mucus also provides protection by capturing AMPs, which act to prevent the pathogens from reaching the epithelium. In the model of Muc2-deficient mice (Muc2 is the gene coding for one of the proteins making up the mucus), an increase in the translocation of commensal bacteria is observed and these animals develop intestinal inflammatory diseases [3].
CROSSTALK BETWEEN THE GUT MICROBIOTA AND THE INNATE IMMUNE SYSTEM
Among the players of the innate immune system which participate in intestinal homeostasis, antigen-presenting cells (APC), such as the macrophages (Mj) and the DCs have a major role. The Mj and the DCs synthesise IL-10 and thus promote differentiation of Treg [4] and the maturation of the Th17 lymphocytes via the implication of commensal bacteria: the segmented filamentous bacteria (SFB). These have the particular ability to adhere to the intestinal epithelial cells causing active stimulation of the immune system [5] (Figure 3). A study shows that colonisation of mice by these SFB, induces the differentiation of Th17 thus resulting in protection from Citrobacter rodentium (the murine equivalent of EPEC and EHEC). It has been suggested that this protection is due to the capacity of the SFB to cause Th17 to stimulate the synthesis of IL-22, a cytokine known to stimulate the synthesis of AMPs [6]. To come back to the DCs, these, by extending their dendrites between the epithelial cells, are able to phagocyte the bacteria present in the intestinal lumen. These commensal bacteria are then transported to the mesenteric lymph nodes to induce the production of IgA secreted by the plasma cells [1]. The innate lymphoid cells (ILC) also play an important role in intestinal homoeostasis; this is related to their capacity to initiate and direct intestinal immune responses. More specifically, the type 3 ILCs (ILC3) have a unique place in the interaction with the gut microbiota. By synthesising IL-22, these cells stimulate the production of mucus, AMPs and the secretion of chemokines and recruitment of polymorphonuclear (PMN) cells (Figure 2) [1].
CROSSTALK BETWEEN THE MICROBIOTA AND THE ADAPTIVE IMMUNE SYSTEM
The final maturation of the adaptive immune system is characterised by the colonisation of the intestinal mucosa by mature effector T-lymphocytes with inflammatory properties (Th17), T-lymphocytes with antiinflammatory properties (Treg) and B-lymphocytes (Figure 2). Besides effects on the macrophages and the differentiation of the Th17 cells, the SFB also stimulate the development of the lymphoid follicles and participate in the differentiation of the B-lymphocytes to IgA-producing plasma cells the action of which is the containment of pathogenic bacteria in the mucus [5]. Other commensal bacteria can stimulate adaptive immune responses: a mixture of 17 Clostridia strains isolated from a human faecal sample and introduced in mice induced an anti-inflammatory response by stimulating the Treg [7]. Faecalibacterium prausnitzii has also been identified for its anti-inflammatory action in vitro and in vivo by acting on the NF-kB factor, DCs and Mj which secrete IL-10 and enhance differentiation of Treg to the detriment of Th17 [8]. Of the Bacteroidetes, Bacteroides fragilis and B. thetaiotaomicron have also been described as exerting anti-inflammatory activity. B fragilis synthetises a polysaccharide A (PSA) that prevents pro-inflammatory IL-17 production and stimulates the anti-inflammatory of secretion IL-10 (Figure 3). In a specific model of Helicobacter hepaticus-induced colitis, PSA stimulated the development of lymphoid follicles, stimulated Treg lymphocyte cells and protected the mice [9].
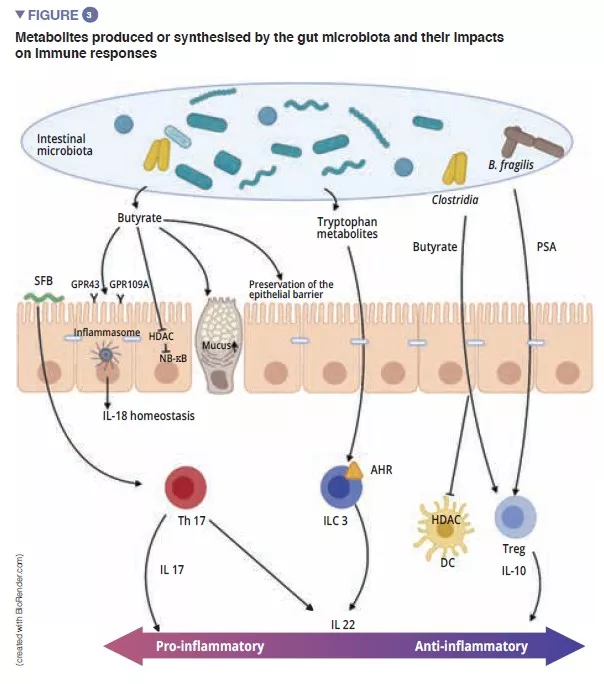
MICROBIAL METABOLITES: IMPORTANT MEDIATORS IN THE CROSSTALK BETWEEN THE MICROBIOTA AND ADAPTIVE IMMUNITY
Short-chain fatty acids (SCFAs), tryptophan metabolites and bile salts are the principal metabolites produced by the gut microbiota which exert a protective effect against infections [9, 10]. Butyrate, propionate and succinate are known to act on intestinal homoeostasis, on mucus secretion, but also on the various cells of the immune system. Among other effects, butyrate has anti-inflammatory and anti- microbial effects. This action is exerted via the G-coupled protein receptors (GPR) found on the epithelial cells and the macrophages [9]. F prausnitzii produces large quantities of butyrate, which may partly explain its anti-inflammatory effect. It inactivates NF-kB and thus suppresses synthesis of the pro-inflammatory cytokines IFN-g, TNF-a, IL-1b, IL-8 by the enterocytes [8] (Figure 3) . It also induces metabolic and epigenetic modifications (via histone deacetylases, HDACs) macrophages in mice, thus amplifying their anti-microbial activities in vitro and in vivo [11]. Commensal bacteria can also metabolise tryptophan and produce antimicrobial substances. An example is the Lactobacilli, which utilise it as an energy source to synthesise an indole that binds to aryl hydrocarbon receptors (AhR) present on the ILC3. AhR triggers IL-22 secretion by the ILCs and this further drives the secretion of AMPs and protects against infections [9].
MICROBIOTA – INTESTINAL IMMUNE SYSTEM CROSSTALK FOR PROTECTION AGAINST VIRAL INFECTIONS
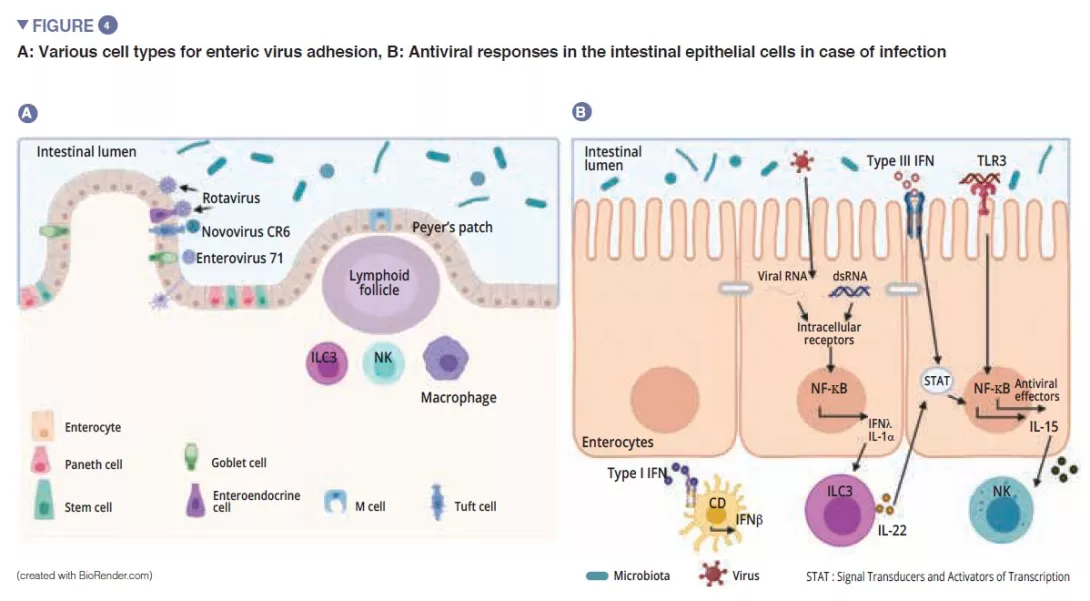
Among the enteric viruses, norovirus and rotavirus are the main causes of gastroenteritis [12]. The enteric viruses infect various cell types: enterovirus 71 specifically infect the goblet cells, whereas the rotavirus has a preferential tropism for the enterocytes [13] (Figure 4A). The gut microbiota acts as a barrier against enteric viral infections. The viruses have evolved and become adapted to their host, implementing mechanisms that enable them to cross the intestinal barrier and escape barrier immunity: it is in fact difficult to infect mice effectively with human enteric viruses by the oral route [13]. Virus penetration into the enterocyte triggers the secretion of type III interferon (IFN). Detection of a virus can induce IL-la, which activates the ILC3 to produce IL-22. This IL protects against enteric viral infections and acts synergistically with type III IFN to induce the expression of antiviral effectors and IL-15. Recognition of a virus by TLR-3 leads to the activation of the NF-kB pathway and to the production of IL-15 also. IL-15 activates the cytotoxic lymphocytes (NK cells). Those viruses, which have traversed the intestinal barrier, trigger the production of type I IFN by the macrophages of the lamina propria (Figure 4B). Some enteric viruses (rotavirus, reovirus, enterovirus) are able to adhere to the intestinal bacteria, enhancing penetration into the intestinal epithelial cells [13]. The SFB, which accelerate epithelial cell turnover produce protection against rotavirus infection in mice by expulsing infected cells [14]. The bile acids metabolised by the gut microbiota also act to protect the small intestine (but not the colon) from acute infection by norovirus in mice by enhancing the production of type III IFN in the small intestine [15].
Conclusion
The study of the relationship between the gut microbiota and intestinal immune response represents significant progress in gastroenterology research. Intestinal homeostasis is maintained due to the recognition of commensal bacteria by the cells of the innate system and the cells of the intestinal epithelium, either by direct contact (in the case of SFB), or via the synthesis of metabolites by the microbiota. The loss of homeostasis (intestinal dysbiosis, infections etc.) causes stimulation of the innate responses and an activation of the adaptive system. Poor “management” of inflammation can result in the onset of disease, such as post infectious irritable bowel syndrome.
1 Perez-Lopez A, Behnsen J, Nuccio SP, Raffatellu M. Mucosal immunity to pathogenic intestinal bacteria. Nat Rev Immunol 2016 ; 16 : 135-48.
2 Allaire JM, Crowley SM, Law HT, et al. The intestinal epithelium: central coordinator of mucosal immunity. Trends Immunol 2018 ; 39 : 677-96.
3 Van der Sluis M, De Koning BA, De Bruijn AC, et al. Muc2-deficient mice spontaneously develop colitis, indicating that MUC2 is critical for colonic protection. Gastroenterology 2006 ; 131 : 117-29.
4 Kim M, Hill A A, Wu WJ, et al. Intestinal microbes direct CX3CR1+ cells to balance intestinal immunity. Gut Microbes 2018 ; 17 : 151-63.
5 Flanningan KL, Denning TL. Segmented filamentous bacteria-induced immune responses: a balancing act between host protection and autoimmunity. Immunology 2018 ; 154 : 537-46.
6 Ivanov I I, Atarashi K, Manel N, et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 2009 ; 139 : 485-98.
7 Atarashi K, Tanoue T, Oshima K, et al. Treg induction by rationally selected mixture of Clostridia strains from the human microbiota. Nature 2013 ; 500 : 232-6.
8 Miquel S, Martin R, Rossi O, et al. Faecalibacterium prausnitzi and human intestinal health. Curr Opin Microbiol 2013 ; 16 : 255- 61.
9 Levy M, Blacher E, Elinav E. Microbiome, metabolites and host immunity. Curr Opin Microbiol 2017 ; 35 : 8-15.
10 Michaudel C, Sokol H. The gut microbiota at the service of immunometabolism. Cell Metabolism 2020 ; 32 : 514-23.
11 Schulthess J, Pandey S, Capitani S, et al. The short Chain Fatty Acid Butyrate imprints an antimicrobial program in macrophages. Immunity 2019 ; 50 : 432-45.
12 Bányai K, Estes MK, Martella V, et al. Viral gastroenteritis. Lancet 2018 ; 392 : 175-86.
13 Segrist E, Cherry S. Using diverse model systems to define intestinal epithelial defenses to enteric viral infections. Cell Host Microbe 2020 ; 27 : 329-44.
14 Shi Z, Zou J, Zhang Z, et al. Segmented filamentous bacteria prevent and cure rotavirus infection. Cell 2019 ; 179 : 644-658.e13.
15 Grau KR, Zhu S, Peterson ST, et al. The intestinal regionalization of acute norovirus infection is regulated by the microbiota via bile acid-mediated priming of type III interferon. Nat Microbiol 2020 ; 5 : 84-92.