Resilience of healthy adult gut microbiota following antibiotic exposure
Commented articles - Adults' section
By Pr. Harry Sokol
Gastroenterology and Nutrition, Saint-Antoine Hospital, Paris, France
Lay public section
Find here your dedicated section
Sources
This article is based on scientific information
Sections

About this article
Author
Comments of the original article of Palleja et al. (Nature Microbiology 2018)
To minimize the impact of antibiotics, gut microorganisms harbour and exchange antibiotics resistance genes, collectively called their resistome. Using shotgun sequencing-based metagenomics, we analysed the partial eradication and subsequent regrowth of the gut microbiota in 12 healthy men over a 6-month period following a 4-day intervention with a cocktail of 3 last-resort antibiotics: meropenem, gentamicin and vancomycin. Initial changes included blooms of enterobacteria and other pathobionts, such as Enterococcus faecalis and Fusobacterium nucleatum, and the depletion of Bifidobacterium species and butyrate producers. The gut microbiota of the subjects recovered to near-baseline composition within 1.5 months, although 9 common species, which were present in all subjects before the treatment, remained undetectable in most of the subjects after 180 days. Species that harbour β-lactam resistance genes were positively selected for during and after the intervention. Harbouring glycopeptide or aminoglycoside resistance genes increased the odds of de novo colonization, however, the former also decreased the odds of survival. Compositional changes under antibiotic intervention in vivo matched results from in vitro susceptibility tests. Despite a mild yet long-lasting imprint following antibiotics exposure, the gut microbiota of healthy young adults are resilient to a short-term broad-spectrum antibiotics intervention and their antibiotics resistance gene carriage modulates their recovery processes.[1]
What is already known about this topic?
The human gut microbiota forms a complex and balanced ecosystem. Perturbations of this ecosystem can play a role in the development of infections, obesity, diabetes as well as neurological and inflammatory disorders. It is estimated that antibiotics have added 2 to 10 years to our life expectancy, but early exposure to these drugs has also been associated with noxious metabolic, inflammatory and neurological effects, both in animal models and in humans. When microbial communities are exposed to antibiotics, not only do they react by shifting their composition, but also by evolving, optimizing and disseminating antibiotic resistant genes (ABR genes) which collectively form the resistome.[2] The human gut microbiota is a reservoir of ABR genes which are exchanged between the resident strains, thereby propagating resistance.[3] The development and spread of antibiotic-resistant bacteria constitute a serious threat to public health. Only a few studies have investigated the effects of specific antibiotics on intestinal ecosystems and their associated resistomes. In previous work it was shown that antibiotic administration induces a decrease in microbiota diversity and an in- crease in the carriage of ABR genes.[4, 5] However, the effects of a combination of antibiotics on the microbiota and the role of ABR genes in microbial persistence have not yet been studied. In this study, 12 healthy men aged 18 to 40 years received a cocktail of three last-resort antibiotics (vancomycin, gentamicin and meropenem). The authors analysed the impact of this treatment on the gut microbiota by shotgun sequencing of faecal samples taken before and at four time points over a 6-month period following antibiotic administration.
Cornerstone of the modern therapeutic arsenal, antibiotics saved millions of lives. On the other hand, their excessive and sometimes inappropriate use can lead to the emergence of multiple forms of resistance in microorganisms. Each year, the World Health Organization (WHO) organizes the World AMR Awareness Week (WAAW) to increase awareness of this public health issue. Read the dedicated page:
Microbiota at the forefront of antibiotic resistance
What is the World AMR Awareness Week?
Each year, since 2015, the WHO organizes the World AMR Awareness Week (WAAW), which aims to increase awareness of global antimicrobial resistance. Held on 18-24 November, this campaign encourages the general public, healthcare professionals and decision-makers to use antimicrobials carefully, to prevent the further emergence of antimicrobial resistance.
What are the main results of this study?
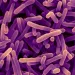
At D4, immediately after the intervention, microbiota diversity and richness were notably reduced compared to baseline. However, despite the use of very broad spectrum antibiotics, many species were still detectable at D4 (Figure 1a). By D8, microbiota diversity (measured by the Shan- non index) had considerably increased, suggesting that surviving microorganisms had begun to regrow (Figure 1b). At 6 months, microbiota diversity was almost completely restored to baseline levels but this was not the case for richness, suggesting that some strains had been permanently (or at least extendedly) eradicated.
Some of the early observable changes included blooms of normally subdominant commensals like Escherichia coli, Veillo- nella spp., Klebsiella spp., Enterococcus faecalis and Fusobacterium nucleatum and a sharp depletion of butyrate-producers like Faecalibacterium prausnitzii, Roseburia hominis, Anaerostipes hadrus, Coprococcus spp. and Eubacterium spp. These shifts in microbiota composition were no longer significant at D42.
The authors then investigated the role of ABR genes in microbiota recovery. In particular, they found that β-lactamase-harbouring metagenomic species had significantly higher odds of survival (OR = 1.64 [1.24-2.17]) at D8. In addition, metagenomic species not detected at baseline had better odds of subsequent de novo colonization if they harboured ABR genes against one of the three antibiotic classes used.
Key points
-
The intestinal microbiota of healthy young adults is resilient to 4 days of broad-spectrum antibiotic treatment with an approximate 6-month recovery of most bacterial communities.
-
The recovery of individual species is modulated by ABR gene carriage.
-
The impact of prolonged or repetitive antibiotic treatment requires further study, especially in paediatric populations.
What are the practical consequences?
These findings indicate that the gut micro- biota of healthy young adults is resilient to a 4-day intervention with broad spectrum antibiotics with recovery of the majority of bacterial communities after about 6 months. The recovery of individual species is modulated by carriage of ABR genes. Further studies are needed to assess the impact of repetitive perturbations and/or over longer periods and to determine whether these findings also hold true in children whose immune system and microbiota are immature. It is possible that repetitive use of antibiotics over long periods selects for bacteria carrying ABR genes at the expense of other commensals, with prolonged or permanent effects on the microbiota. In such cases, corrective intervention with exogenous supply of microorganisms could be considered. The effects of antibiotics on the intestinal microbiota are therefore important and their use must be rationalized.
Conclusion
Broad spectrum antibiotics negatively impact the gut microbiota in an immediate, significant and durable manner for some species. In healthy young adults, the gut microbiota is resilient but near-complete recovery takes about six months. Modulated by ABR gene carriage, the capacity of species to regenerate is more favourable to diversity than to richness.