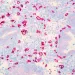
A gene catalog for the vaginal microbiota
A catalog of all vaginal bacterial species and their gene function is now available. This public resource will facilitate the work of researchers and provide a better understanding of the role played by vaginal microorganisms in women’s health.
Sources
This article is based on scientific information
About this article
By 2003, the complete human genome had been decoded. In 2008, genome decoding was launched for human microbial populations (MetaHIT and (sidenote: Human Microbiome Project ) projects), followed by an analysis of their health roles ( (sidenote: International Human Microbiome Consortium ) ). While these databases are essential for understanding the structure and function of microbial communities and their role in diseases, they mainly focus on the gut microbiota. This situation changed in early 2020 with the publication of (sidenote: Vaginal integrated non-redundant gene catalog ) , a gene catalog for the vaginal microbiota.
VIRGO: the most complete vaginal gene catalog
VIRGO was built using metagenomic data (n = 264) and data from complete genomes (n = 308) gathered through urogenital samples and isolates. For the moment, the database mainly relates to bacteria, containing only a few viral and fungal gene sequences. It catalogs almost a million (sidenote: That do not create duplicate code for the same protein ) bacterial genes, classifying them both by taxonomic rank and function. VIRGO covers more than 95% of human vaginal microbiota and can be used for (sidenote: North America, Africa and Asia ) , making it possible to characterize genes and analyze their abundance and expression in the vaginal microenvironment. It has already shown that there is much more intra-species diversity within the vagina than previously thought, to the point of calling into question the idea that a major strain of Lactobacillus is dominant.
VOG: protein families grouped by function
The genes identified in VIRGO were then translated and grouped by protein family and function in a second catalog, VOG (Vaginal Orthologous Groups). The team used this catalog to search for new protein variants. The result was the identification of a previously unknown alanine-to-valine substitution in the protein sequence of a toxin excreted by Gardnerella vaginalis. The catalog will therefore make it possible to identify different variants of the same protein, suggest a biological significance for these variants and put forward new hypotheses.
A fast and accurate tool
VIRGO is a fast, accurate and multi-faceted tool for characterizing vaginal microbiota: it provides a global view of vaginal bacteria groups and has a gene-centric design that allows characterization by function and taxonomy; it also has high scalability, high sensitivity–making it possible to characterize low-abundant bacteria–, and is a simple tool for assessing genetic richness and intra-species diversity. It will be particularly valuable for users with limited computer skills, a large volume of data to sequence, and/or a limited computer infrastructure.