Cholesterol metabolism by uncultured human gut bacteria influences host cholesterol level
Commented articles - Adults' section
By Pr. Harry Sokol
Gastroenterology and Nutrition Department, Saint-Antoine Hospital, Paris, France
Lay public section
Find here your dedicated section
Sources
This article is based on scientific information
Sections

About this article
Author
Comments on the original article of Kenny et al. (Cell Host & Microbe 2020)
The human microbiome possesses extensive metabolic capabilities but our understanding of the mechanisms linking gut microbes to human metabolism remains limited. In this article, the authors focused on the conversion of cholesterol to the poorly absorbed sterol coprostanol by the gut microbiota to develop a framework for the identification of functional enzymes and microbes. By integrating paired metagenomics and metabolomics data from e xisting cohorts with biochemical knowledge and experimentation, the authors predicted and validated a group of microbial cholesterol dehydrogenases that contribute to coprostanol formation. These enzymes are encoded by ismA genes in a group of uncultured microbes, which are prevalent in geographically diverse human cohorts. Individuals harbouring coprostanol-forming microbes have significantly lower faecal cholesterol levels and lower total serum cholesterol with effects comparable to those attributed to variations in lipid homoeostasis genes. Thus, cholesterol metabolism by these microbes may play important roles in reducing intestinal and serum cholesterol concentrations, directly impacting human health. [1]
What do we already know about this subject?
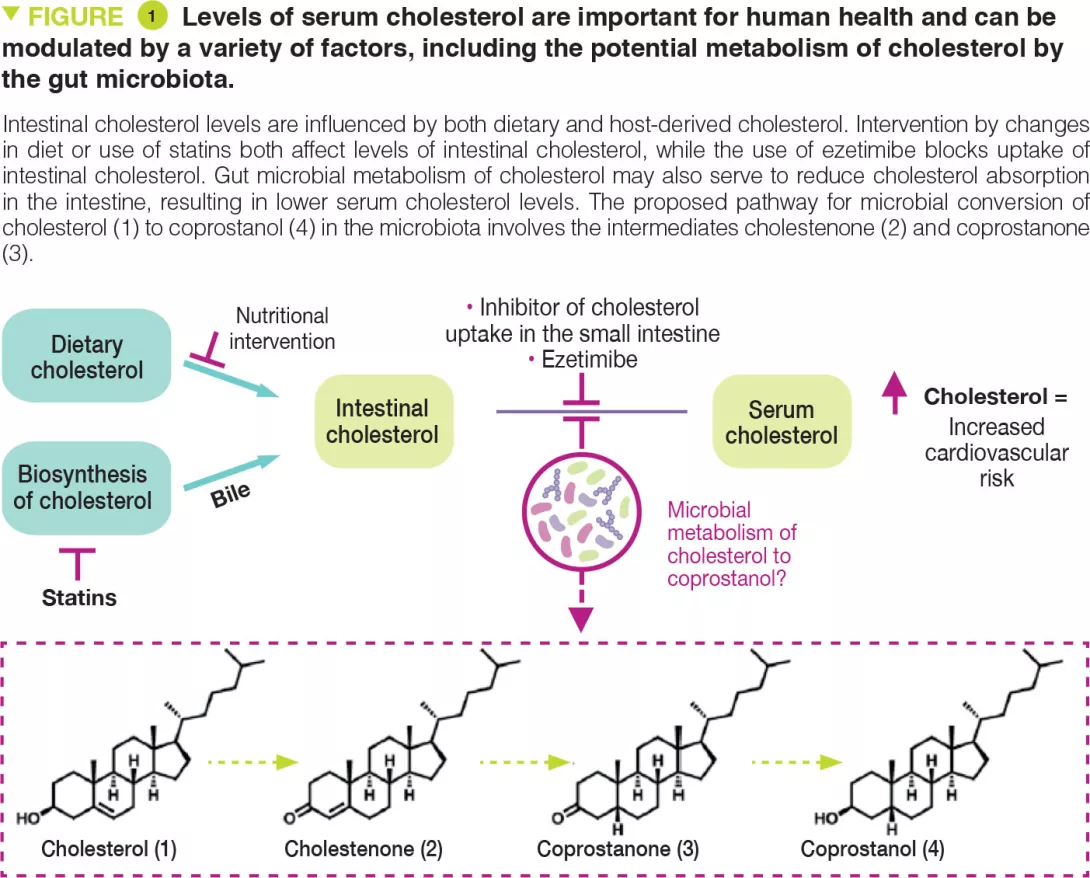
Cholesterol is a key biological molecule that functions as a structural component of all animal cell membranes and is a pre-cursor of steroid hormones, vitamin D, and bile acids. Two main sources of cholesterol are thought to influence concentrations of this metabolite in serum: endogenous cholesterol synthesised in the liver and exogenous cholesterol derived from dietary components of animal origin (Figure 1). The cholesterol synthesised in hepatocytes is transported to the gallbladder and is then secreted into the small intestine along with other bile salts. In the intestine, biliary cholesterol (~1–2 g/day) mixes with dietary cholesterol (~0.2–0.4 g/day in the average American diet), and both sources are eventually transported into enterocytes for packaging into lipoprotein particles and secretion into the plasma. Hypercholesterolaemia is a risk factor for cardiovascular disease (CVD), which is the cause of one-fourth of all deaths in industrialised countries.
Reducing cholesterol transport in the intestine is a clinically validated strategy for lowering serum cholesterol levels. A range of gut microbes metabolise and modify dietary and host-derived molecules in the small intestine. Because both sources of cholesterol pass through this environment, the gut microbiota may influence serum cholesterol levels. Indeed, microbiota transfer from human donors with elevated serum cholesterol levels can impart this hypercholesterolaemia phenotype to mice. [2, 3] Other studies have reported that administering specific bacterial species can have cholesterol-lowering effects. [4] However, the precise mechanisms underlying these observations are currently unknown. The gut microbiota may exert cholesterol-lowering effects by metabolising intestinal cholesterol to coprostanol (Figure 1), which would reduce the amount of cholesterol absorbed from the intestine.
This microbiota-dependent transformation has been known to occur in humans since the early 1900s. Several coprostanol-generating gut bacteria with similar physical and biochemical characteristics have been reported from a variety of different sources including rats, baboons, and humans. However, most of these strains are not currently available and were never sequenced. Early work showed that coprostanol formation by this group of gut bacteria proceeds through an indirect reduction pathway involving the initial oxidation of cholesterol (1) to cholestenone (2), followed by reduction of the D4,5 double bond to form coprostanone (3), and subsequent re-reduction of ketone to generate coprostanol (4) (Figure 1). The bacterial enzymes responsible for this metabolism were never identified. More recently, other reports have implicated additional phylogenetically diverse gut bacteria in coprostanol formation. [5] While efforts to elucidate how gut microbial metabolism of cholesterol affects human serum cholesterol levels span over 100 years, mechanistic support for this connection has remained elusive due to a limited understanding of the gut microbes, genes, and enzymes responsible for coprostanol formation.
Key points
-
Some bacteria found in the human intestinal microbiota possess enzymes belonging to the ismA family capable of breaking down cholesterol.
-
The presence of ismA+ species in gut microbiota is associated with decreased faecal and serum cholesterol in humans.
-
The effect of ismA+ species on serum cholesterol is on par with human genetics.
What are the main insights from this study?
The authors used a multi-disciplinary strategy to discover gut bacterial enzymes. This strategy, based on correlations between metagenomics and metabolomics data from existing human cohorts, identified and characterised an extensive family of cholesterol dehydrogenase enzymes from a clade of uncultured intestinal bacteria implicated in the metabolism of cholesterol to coprostanol. Firstly, the enzyme responsible for the first step in cholesterol transformation, called ismA, was identified in Eubacterium coprostanoligenes, a bacteria already known for this function. Analysis of sequencing data from human cohorts then identified homologous enzymes in a group of uncultured anaerobic bacteria.
The presence of these ismA genes in the microbiome was associated with the presence of coprostanol in stools and lower faecal cholesterol levels. Finally, to demonstrate the potential for these cholesterol- metabolising bacteria to influence human health, the authors showed that presence of ismA genes in human metagenomes is associated with a decrease in total cholesterol concentrations in serum that is on par with the effects observed from variants in human genes involved in lipid homoeostasis
What are the consequences in practice?
Overall, these findings confirm the role of gut-bacterial metabolism in modulating host cholesterol levels in the intestine and also, more importantly, on a systemic level. This work paves the way for the use of the gut microbiota as a predictive biomarker of high cholesterol and establishes the foundations for microbiota-targeted therapeutic interventions.
Conclusion
This study highlights the role of the gut microbiota in breaking down cholesterol with an effect on serum cholesterol levels. Gut microbiota could soon become the target of cholesterol lowering therapies.