SCFAs: new intestinal markers of Parkinson’s disease?
Short-chain fatty acids (SCFAs) are gut microbiota metabolites already known to be involved in Parkinson’s disease, particularly in rodents. But what about in humans?


Short-chain fatty acids (SCFAs) are gut microbiota metabolites already known to be involved in Parkinson’s disease, particularly in rodents. But what about in humans?

(sidenote: https://parkinsonscare.org.uk/worldparkinsonsday/ ) (April 11) is an opportunity each year for patient associations and health professionals to see how research is progressing on this complex neurodegenerative disease for which there is no treatment to date. This year, discussions will no doubt focus on a new study by a team of Taiwanese researchers. The team wanted to determine whether SCFAs could discriminate between Parkinson’s patients and healthy individuals, and whether there was a correlation with disease severity. To do this, they analyzed plasma and fecal levels of several subtypes of SCFAs in addition to the gut microbiota of 181 participants (96 patients and 85 controls). They also studied the motor and cognitive effects of the disease. The study results were published in Neurology in early 2022.
In Parkinson’s patients, the results showed reduced fecal SCFA levels compared to healthy individuals (butyric acid, valeric acid and propionic acid), whereas plasma levels were higher.
Another takeaway: fecal quantities of the same SCFAs were lower in patients with advanced Parkinson’s disease compared to patients in the early stages of the disease.
Fecal and plasma concentrations vary depending on the severity of motor and cognitive symptoms.
More severe motor impairment is correlated with a low fecal concentration of most SCFAs, coupled with an increased plasma propionic acid concentration.
More serious cognitive symptoms are associated with lower fecal levels of butyric acid and higher plasma concentrations of butyric acid and valeric acid.
The study showed that microbial diversity in Parkinson’s patients differed from that in healthy patients. This study highlights the correlation between SCFA concentrations and the abundance of pro-inflammatory bacteria (Clostridiales and Ruminococcus), particularly in the case of propionic acid. This supports the hypothesis that gut inflammation is associated positively with disease aggravation.
Specifically, these results suggest that SCFAs may serve as gut biomarkers for Parkinson’s disease and also reflect severity. This is an interesting scientific advance, which may pave the way for more accurate diagnosis and better patient management.
By Pr. Gianluca Ianiro
Gastroenterology Department, Fondazione Policlinico Universitario Agostino Gemelli-IRCCS, Università Cattolica del Sacro Cuore, Rome, Italy.

Recently, a new viral challenge has appeared on social media, including Twitter and Instagram: the #BluePoopChallenge. This challenge consists in eating blue-colored food (specifically, two muffins made with royal blue food dye) and record how long does it take to see blue stools in your toilet.
As for all patient-driven examinations, I would first recommend a visit with a health professional with expertise in digestive disease to avoid the do it yourself approaches

This method was recently shown to be a simple and direct way to measure the gut transit time. It means how long the food takes to pass throughout our gastrointestinal tract, from eating to evacuation, and it is usually assessed to evaluate gut motility, which is a key component of gut health. In a recent study [1] by Asnicar and colleagues, published in Gut, authors evaluated the blue dye method as a marker of gut transit time, and its association with specific health markers (including stool consistency and frequency, gut microbiome composition and function, and cardiometabolic health) in 863 healthy individuals.
First, they found that harder stool consistency, measured with British Stool Chart, is associated with a longer gut transit time (>5 days median for type 1), while softer consistency corresponds to shorter gut transit time (1 day median for type 6).
Moreover, they also found that gut transit time is associated with several features of gut microbiome. Alpha diversity, which is a marker of microbial health, was positively correlated with gut transit time. Longer transit time was associated with specific microbial taxa, including Akkermansia muciniphila (a beneficial strain with favorable metabolic properties), Bacteroides spp. and Alistipes spp. Generally, gut transit time was associated with gut microbiome features more than stool consistency or stool frequency.
Finally, longer transit time was associated with visceral fat mass and postprandial lipid and glucose responses (both cardiovascular risk factors).
In conclusion, blue dye method appears to be a simple and inexpensive marker of gut transit time, which was found to be associated with markers of human health, including gut microbiome diversity and composition, and specific cardiovascular risk factors. It could be a reliable method to evaluate gut transit time when needed, i.e. in patients with constipation). The way this procedure is being diffused (viral expansion through social media) suggests that lots of patients will require this testing to physicians. As for all patient- driven examinations, I would first recommend a visit with a health professional with expertise in digestive disease to avoid the DYI approaches. Then, if the physicians confirm the need of a gut transit time test, then the blue dye method would be a reliable and cheap one.
Future evaluations, including comparisons with other methods of gut transit assessment, are needed to confirm these results and allow a large-scale positioning of this tool into the diagnostic armamentarium of GI and nutritional disorders.
By Pr. Markku Voutilainen
Turku University Faculty of Medicine; Turku University Hospital, Department of Gastroenterology, Turku, Finland

Predicting the risk of preterm birth using a simple, rapid, and inexpensive method is a challenge for obstetricians, who still lack a reliable predictive method for this clinical complication, the leading cause of death in children under the age of five. The risk factors are well known: vaginal dysbiosis associated with local inflammation. A team of researchers had the idea of using their recently-described DESI-MS (Desorption Electrospray Ionization Mass Spectrometry) analysis method to identify – in less than 3 minutes and without the need for sample preparation – the metabolites present in the cervicovaginal mucosa. Their hypothesi,s? The metabolome thus characterized may make it possible to predict the composition of the vaginal microbiota and local immune and inflammatory responses, and to monitor their development towards states associated with the risk of preterm birth. DESI-MS was used to analyze more than 1,000 cervicovaginal samples from 365 pregnant women in two cohorts.
Among the metabolites detected, 113 made it possible to effectively distinguish between two types of microbiota: one depleted and the other dominated by lactobacilli, a marker of good vaginal health. The metabolic profile obtained using DESI- MS also predicted the levels of several immune markers (IL-1β, IL-8, C3b/iC3b, IgG3, IgG2, MBL – Mannose-Binding Lectine) measured in a subgroup of 391 women. Some of these (C3b, IL-1β, IgG2, IgG3) were found at high levels in Lactobacillus- depleted vaginal microbiomes, indicating activation of the local innate and adaptive immune response. In a final series of tests, the metabolic vaginal profile obtained using DESI-MS could not reliably predict the direct risk of preterm birth. However, the researchers foresee potential clinical applications. Monitoring vaginal metabolites using DESI- MS could help detect changes in the vaginal microbiota and local immune markers associated with preterm birth.
By Pr. Markku Voutilainen
Turku University Faculty of Medicine; Turku University Hospital, Department of Gastroenterology, Turku, Finland
Prostate cancer (PC) is one of the most common cancers in males. Because tumors growth and progression depend on androgen levels, androgen deprivation therapy (ADT), surgical or chemical castration is used to treat patients with PC. However, some of them develop a castration- resistant prostate cancer (CRPC) which result in tumor progression and new treatment strategy are under investigation. Since recent studies have highlighted the role of microbiota in both cancer development and therapy success, the authors used PC mouse models and patient data to examine the role of gut microbiota in PC carcinogenesis. Enrichment of Ruminococcus spp. and Bacteroides acidifaciens was detected after development of CRPC but gut microbiota ablation slowed tumor growth in CRPC mice. Castration resistant (CR) fecal microbiota transplantation (FMT) from castration-resistant (CR) mice and R. gnavus administration led to increased circulating androgen levels and increased PC growth and CRPC development. PC growth was controlled by FMT from hormone sensitive PC individuals and Prevotella stercorea administration. CRPC patients had enrichment of Ruminococcus and Bacteroides genera associatied with poor outcome while hormone- sensitive PC patients had higher abundance of Prevotella genus linked with more favorable outcome.
Commensal gut microbiota in androgen- deprived patients and mice produce androgens that promote PC growth and the development of CRPC via systemic circulation. Modulation of gut microbiota could theoretically be used as additional therapy for PC.
Chronic obstructive pulmonary disease (COPD) refers to lung diseases (emphysema, bronchitis and asthma) characterized by progressive respiratory distress. Recent studies have revealed changes in the gut microbiota linked to disease development in the lungs. While primarily considered a respiratory disease, COPD commonly co-occurs with chronic gastrointestinal tract diseases. In the present study, the authors took an interest in the the gut-lung axis linked to COPD. Stool analyses have revealed that patients with severe COPD had lower abundance of Bacteroidetes but higher abundance of Firmicutes. Of bacterial families, Prevotellaceae abundance was higher in mild COPD, whereas Bacteroidaceae and Fusobacteriaceae abundancies were lower in severe COPD compared to healthy controls. Short chain fatty acid (SCFA) levels were significantly lower in severe COPD. Fecal microbiota transfer (FMT) to mice from COPD patients caused a significant weight reduction and airway mucus hypersecretion in mice. Acceleration of lung function decline was detected in FMT mice during biomass smoke exposure. This study revealed that COPD patients have gut microbiota dysbiosis with reduced SCFA levels. These changes are possibly linked to airway inflammation and COPD progression.
Immune checkpoint inhibitors (ICI) have dramatically improved the prognosis of several advanced cancers. Evidence have shown that gut microbiota may modulate the treatment response for ICI, and may also be involved in the pathogenesis of immune-related adverse events (IRAE). While antibiotics are known to deteriorate the prognosis of ICI treated cancer patients, little is known about the effect on the microbiota of various co-medications when given at ICI initiation. In the present study, the authors examined the effect of co-medications given 1 month before or after administration of ICI therapy on the treatment results and the occurrence of IRAE.
The use of antibiotics, glucocorticoids (daily dose > 10 mg), proton pump inhibitors, psychotropic drugs, morphine and insulin were associated with significantly shortened survival and decreased tumor response. Combination therapy with these drugs decreased survival more than monotherapy. These medications were also associated with decreased incidence of IRAE. Co-administration of statins, angiotensin-converting enzyme inhibitors and/or angiotensin II receptor blockers, non-steroidal anti-inflammatory drugs, aspirin and oral antidiabetic drugs did not impact patient survival. The present study showed that co-medication influences both to the response and IRAE of ICI treatment. The impact of co-medication may be mediated via microbiota or other immunomodulatory mechanisms. In clinical practice, baseline co-medications should be carefully assessed when ICI therapy is planned. Drugs with negative impact on ICI therapy should be avoided when possible.
CONGRESS REVIEW
By Pr. Eamonn M M Quigley
Lynda K and David M Underwood Center for Digestive Disorders, Division of Gastroenterology and Hepatology, Houston Methodist Hospital and Weill Cornell Medical College, Houston, Texas, USA

WCOG 2020 was scheduled to be held in Prague, the capital of the Czech Republic, at the end of 2020 but was postponed due to the Covid pandemic and was delivered as a hybrid meeting from December 9th to 11th 2021. Thanks to the diligence and organizational skills of all involved an outstanding virtual meeting platform was developed which enabled the transmission of symposia that spanned the entire spectrum of the specialty, special key-note lectures, original free papers and posters to the entire world. In these sessions, experts from around the world joined faculty from the Czech Republic to tackle “hot” topics and controversies alike. Here is a focus on the gut microbiome.
In 2020 the theme of World Digestive Health Day, sponsored by WGO, was “Global Issues in the Microbiome”. Eamonn MM Quigley, on behalf of the many colleagues who contributed to this program, presented its proceedings which encapsulated an overview of the gut microbiome. The factors that influence the microbiome and shape its development through that critical and vulnerable period from birth through early childhood were delineated and the role of diet, throughout the lifespan of the individual, stressed. While a role for the microbiome in many disease states has been proposed, many studies describe association, not causation.
One disorder where a role for the microbiome has generated much excitement is irritable bowel syndrome (IBS). Mirjana Rajilic-Stojanovic explored this in detail and delineated the factors that compound the interpretation of microbiome studies in IBS (e.g. small study population size, heterogeneity of phenotype, variable study design) which, no doubt, contribute to the absence of a consistent microbial signature for IBS. She did alert us to two organisms that may be of particular interest: Methanobrevibacter smithii and Faecalibacterium prausnitzii; the former through its production of methane and the latter through its role in signaling to enterochromaffin cells leading to increased biosynthesis of serotonin – a critical enteric nervous system neurotransmitter and target of much IBS pharmacology.
The microbiome offers tremendous therapeutic potential. Francisco Guarner updated the WGO probiotic guideline and the fascinating topic of Fecal Microbiota Transplantation (FMT) was tackled by Pavel Drastich. He contrasted the efficacy of FMT in Clostridioides difficile related disease to the much more mixed data from other disease states. Among the latter, ulcerative colitis provides the best data (though still rated as of only moderate quality). Here, the interpretation of results is bedeviled by the same factors that confound so many FMT studies: variability in population selection, study endpoints and study protocol. The latter includes such vexing variables as use of fresh vs. frozen stool, route of delivery (naso- jejunal, vs. enema, vs. colonoscopic vs. swallowed capsule), number of treatments (single vs. multiple) and source of material (single donor vs. pooled material from multiple donors). Most fascinating was the glimpse he provided into the future of FMT, predicting an evolution from FMT as we currently know it (i.e. whole stool samples) through the development of specific combinations or consortia of microbes (an area of frantic scientific and commercial activity) to the elaboration of biologically and therapeutically active compounds from microbes. Both the promise and the limitations of FMT in IBS were further developed by Mirjana Rajilic-Stojanovic. Here confounders are vividly displayed: a highly heterogenous phenotype with a variable range and severity of symptoms which fluctuate over time, an incomplete understanding of the role of the microbiome and an, at best, speculative concept of how FMT might act. No wonder that results, to date, are so variable.
Francisco Guarner led off one of the opening sessions of the entire program on a topic that I have retained to the end of this piece – the current situation of microbiota research. This was a typically thoughtful and insightful lecture and most timely given the impact of the Covid-19 epidemic and the global antibiotic resistance crisis; do we need any further reminders of the importance of the microbes that co-habit this planet with us? He returned to the global theme by illustrating the primacy of poor diet in global risk mortality but deftly pointed out that these very diets impoverish not just humans but their microbes and butyrate producers in particular. Low levels of gut microbial diversity have been linked to several disease states with one of the most compelling examples being childhood asthma. Looking to the future, he stressed the inadequacies of a taxonomic approach to the study of the microbiome in disease states and illustrated this point by showing that in normal individuals the composition of the gut microbiome is highly dynamic; up to 90% of strains appear and disappear over time! He called, therefore, for a new approach to the definition of normal vs. abnormal microbiomes (or eubiosis vs. dysbiosis, as some prefer) which emphasized functional competence and ecological stability (incorporating such concepts as resistance to community structural change over time and resilience, an ability to rapidly return to baseline when disrupted, such as by an antibiotic). These are key concepts and reflect the current move away from simple enumeration of microbial numbers, species and strains to the description of functional potential using metagenomics and assays of metabolic products through metabolomics and metatranscriptomics.
Overview
By Pr. Conceição Calhau
NOVA Medical School, New University of Lisbon, Portugal
By Pr. Pedro Povoa
NOVA Medical School, New University of Lisbon, Portugal; Polyvalent Intensive Care Unit, Hospital São Francisco Xavier, CHLO, Lisbon, Portugal; Center for Clinical Epidemiology, OUH Odense, University Hospital, Denmark

For the first time, the gut microbiota diversity is pointed out as a prognostic biomarker of Covid-19 severity. Thus, the microbiota changes as reliable biomarkers in the context of Covid-19 represent a key piece of the disease puzzle, thus highlighting the clinical priority for prevention and possibly new therapeutic strategies. In 2020, the new coronavirus severely affected certain groups of the population, more specifically, the elderly and people with obesity, hypertension, diabetes [1]. Interestingly, publications showed that dysbiosis is a common factor in all these patients [2, 3].
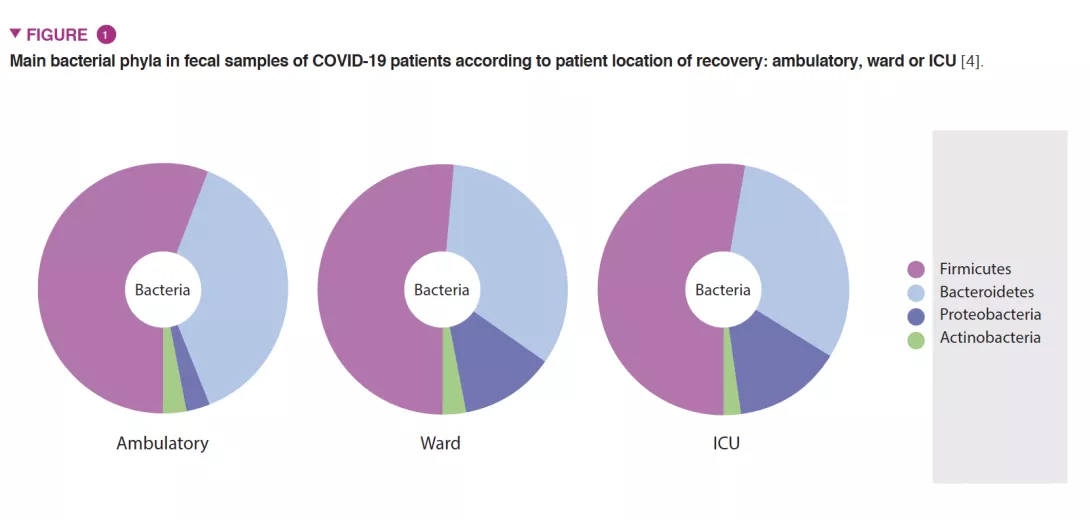
As gut microbiota has been considered a hot topic in the scientific community with a pivotal role on host immune and inflammatory functions, we investigated if changes in gut microbiota composition are associated with an increased clinical severity of Covid-19 [4]. A national multicenter cross-sectional study was conducted in 115 Covid-19 patients categorized by: 1) location of recovery from Covid-19: ambulatory (household isolation), ward or intensive care unit; and 2) Covid-19 severity scale: asymptomatic/mild-to-moderate or severe. Severely ill patients exhibited profound changes in gut microbiota composition compared with mild-to-moderate patients in ambulatory or admitted to the hospital ward (Figure 1). These changes included: 1) lower overall gut microbial diversity; 2) lower abundance of beneficial butyrate-producing bacteria such as Roseburia and Lachnospira; 3) lower Firmicutes/ Bacteroidetes ratio; 4) higher abundance of Proteobacteria. In addition, we detected the virus in fecal samples, which must be considered in public health recommendations [5, 6] Publications from other colleagues have shown that low diversity could be a clinical biomarker predicting greater risk of severity [7-9].
The conventional wisdom was that the healthy lungs were sterile. In the last decade, the application of techniques of microbiota research clearly showed that was not the case. The lungs are colonized by a very low bacterial load compared with the gut [10]. The different parts of the respiratory tract (oropharynx, airway, lungs) present different diversity and compositions related to the colonization sources, colonization rates, extinction rates and distances between each other according to the adapted island model, being the “mainland” the oral cavity [11]. Recent studies in severe Covid-19 patients showed dysbiosis of the airway microbiota (analyzed in BALF samples) similar to the dysbiosis observed during lower respiratory tract infections, pneumonia for example [12, 13]. In addition, Acinetobacter – a common gram-negative non-fermenting bacilli pathogen of ventilator-associated pneumonia that is the most severe ICU-acquired infection in patients undergoing invasive mechanical ventilation – was a common bacterial genus found in lung tissues in deceased patients [14]. The presence of some pathogens in the lung of deceased patients and in the oral cavity is related to the adapted island model migration [15]. As a result of the Covid- 19 associated immune dysregulation, several epidemiologic studies showed an increased risk of hospital acquired infections, namely ventilator associated pneumonia as our group showed. In our study we found that Covid-19 patients presented twice the risk of ventilator associated pneumonia in comparison to non-Covid-19 patients [16].
Microbiota and Covid-19 studies may open perspectives for the development of therapeutic interventions (probiotics, prebiotics…) that aim to correct dysbiosis observed in severe Covid-19 patients. These interventions are expected to increase overall bacterial diversity and the abundance of commensal bacteria, thereby contributing to inhibit the overgrowth of opportunistic pathogens. These studies could also have implications in the design of effective Covid-19 vaccines, as one factor known to control vaccine efficacy could be the gut microbiota.
COMMENTED ARTICLE - CHILDREN’S SECTION
By Pr. Emmanuel Mas
Gastroenterology and Nutrition Department, Children’s Hospital, Toulouse, France

Infants are at a high risk of acquiring fatal infections whose treatment relies on functioning antibiotics. Antibiotic resistance genes (ARGs) are present in high numbers in the gut microbiomes of antibiotic-naive infants, and infant mortality caused by resistant infections is high.
In this article the authors aim to determine the impact of early exposure to formula on the ARG load in neonates and infants born either pre- or full-term. One assumption was that diet exerts selective pressure which influences the microbial community in the gut of infants and formula exposure increases the abundance of taxa-carrying ARGs.
The study showed that formula-fed infants had a higher relative abundance of opportunistic pathogens such as Staphylococcus aureus, S. epidermidis, Klebsiella pneumoniae, K. oxytoca and Clostridioides difficile. Formula-fed infants also had significantly fewer typical infant bacteria such as bifidobacteria, which have potential health benefits.
The novel finding of a correlation between formula exposure and higher neonatal ARG burden shows that clinicians should consider feeding mode in addition to antibiotic use in the first months of life to minimise the proliferation of antibioticresistant gut bacteria in infants.
Antibiotic-resistant bacteria are the cause of many neonatal deaths. The emergence of resistant bacteria is favoured by the use of antibiotics, which is associated with a higher number of antibiotic-resistance genes (ARGs) carried by these resistant or multi-resistant bacterial strains. Mobile genetic elements (MGEs) transmit ARGs between bacteria. It is also known that the type of diet modifies the intestinal microbiota, as well as the quantity of ARGs. The magnitude of the impact of infant diet on resistance has not been adequately described in the literature.
The authors included 46 infants born prematurely between 26 and 37 weeks of gestation of which 21 were fed with formula, 20 with fortified breast milk and 5 with breast milk. Stools were collected within 36 days to analyse the composition of gut microbiota and presence of ARGs. Thirty infants received an antibiotic treatment: stools were collected approximately two weeks after the end of treatment to limit confounding effects.
To compare the results with literature data, a meta-analysis of five studies including 696 neonates with similar data was analysed in parallel.
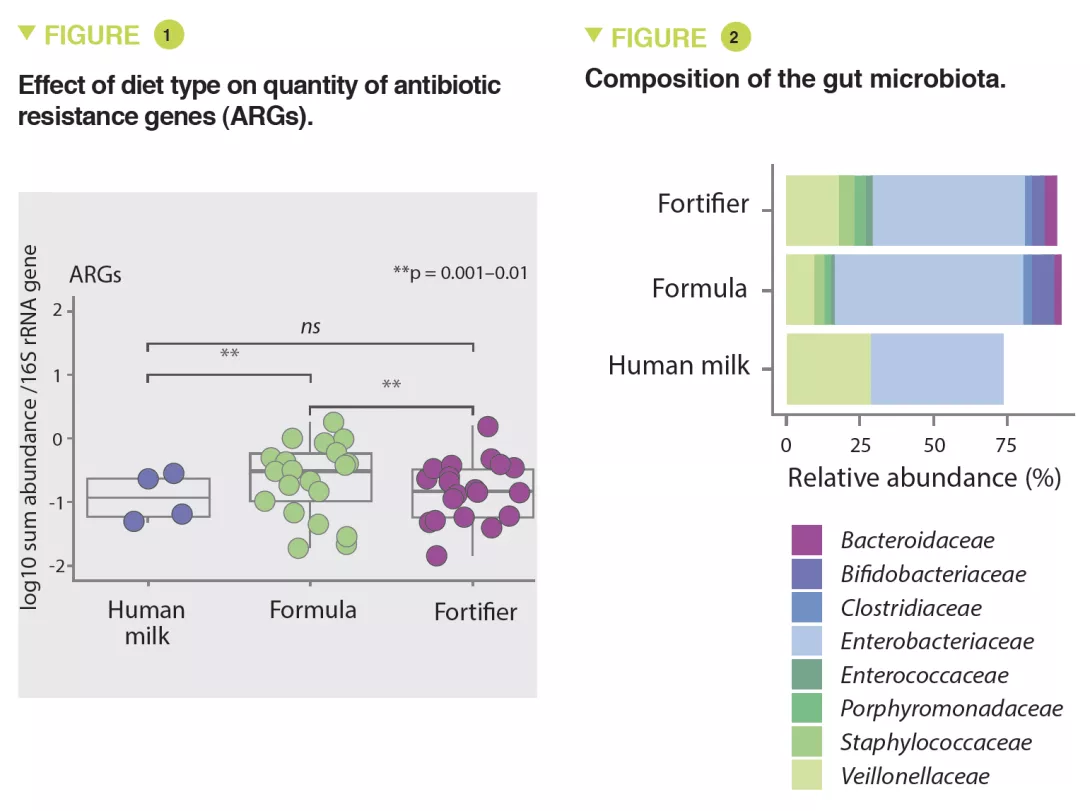
The results showed that infants fed with formula had significantly higher amounts of ARGs than infants fed with fortified breast milk (x 3.6; 95% CI, 1.61-8.9) or breast milk (x 4.3; 95% CI, 1.61-11.56) (p < 0.01) (Figure 1). The abundance of MGEs was similarly increased (p < 0.05).
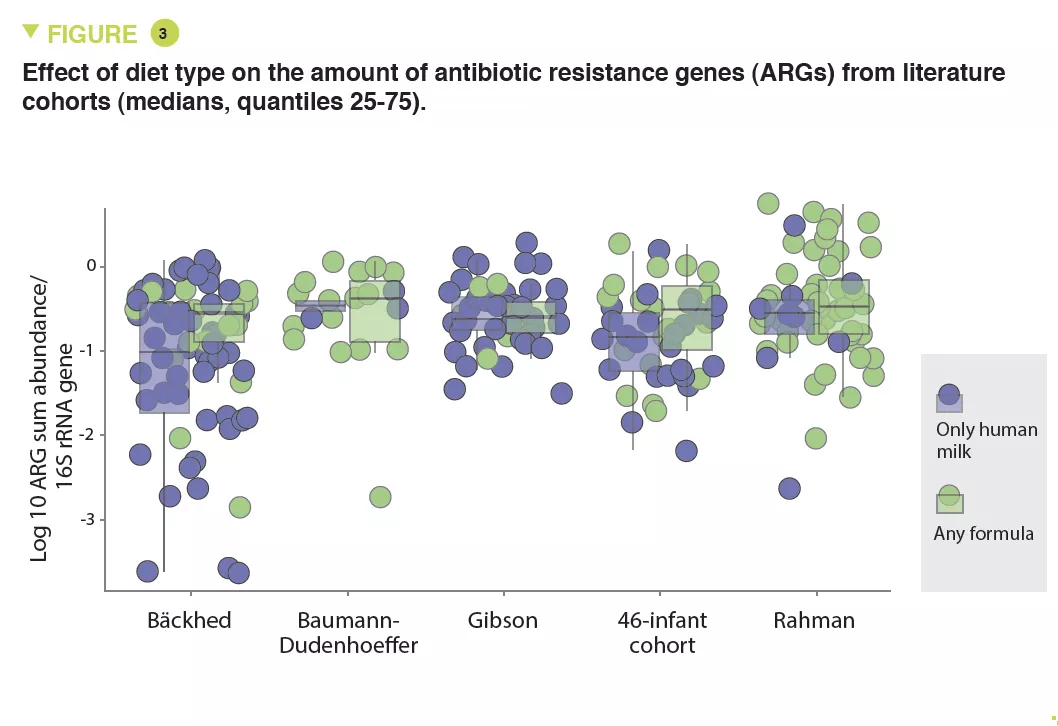
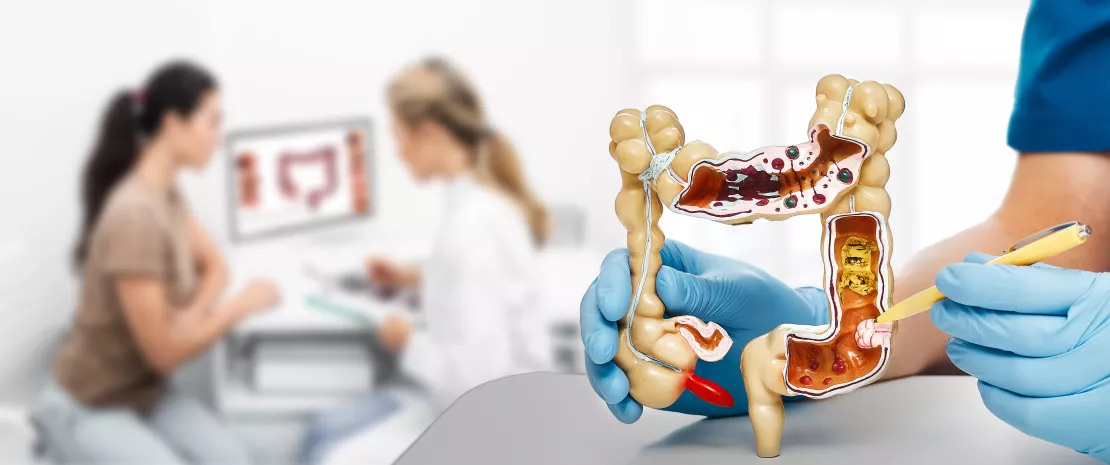
The abundance of Enterobacteriaceae, whose genome is known to contain more mobile ARGs, was higher in formula-fed infants (p < 0.05) (Figure 2) and tended to be inversely correlated to gestational age (p < 0.1). It was noted that the longer the gestation the lower the abundance of these ARGs (x 0.72; 95% CI, 0.57-0.89) (p < 0.001). Several ARGs were significantly more abundant in formula-fed infants including genes encoding the extended-spectrum beta-lactamase present in Klebsiella (p < 0.05). Similar results were observed in the meta-analysis with a 70% relative increase in ARGs in formula-fed neonates (p = 0.013). The median ARG load was higher in the formula-fed infants in all cohorts (Figure 3). Finally, an analysis of gut microbiota revealed that bacteria belonging to Bifidobacteriaceae, Veillonellaceae, Clostridiaceae, Lachnospiraceae, and Porphyromonadaceae families (including strict anaerobe bacteria) were depleted in neonates fed with infant formula; conversely, facultative anaerobe bacteria belonging to the Enterobacteriaceae, Staphyloccoccaceae, and Enterococcaceae families were increased (p < 0.05).
Similarly, several potentially pathogenic species including facultative anaerobe species such as S. aureus, S. epidermidis, K. pneumoniae, K. oxytoca, and a strict anaerobe species Clostridioides difficile, were enriched in formula-fed neonates (p < 0.001). Thus, the use of infant formulas enhances the proliferation of pathogenic bacteria with ARGs.
These results support the benefit of breastfeeding. Feeding premature neonates with infant formula is associated with a 70% increase in ARGs compared to those fed exclusively with breast milk. Enriched breast milk results in a smaller increase in these ARGs.
In addition to the appropriate use of antibiotics, it is important to consider the type of diet for premature neonates, with a preference for breast milk to avoid the proliferation of resistant bacteria.
COMMENTED ARTICLE - ADULTS’ SECTION
By Pr. Harry Sokol
Gastroenterology and Nutrition Department, Saint-Antoine Hospital, Paris, France
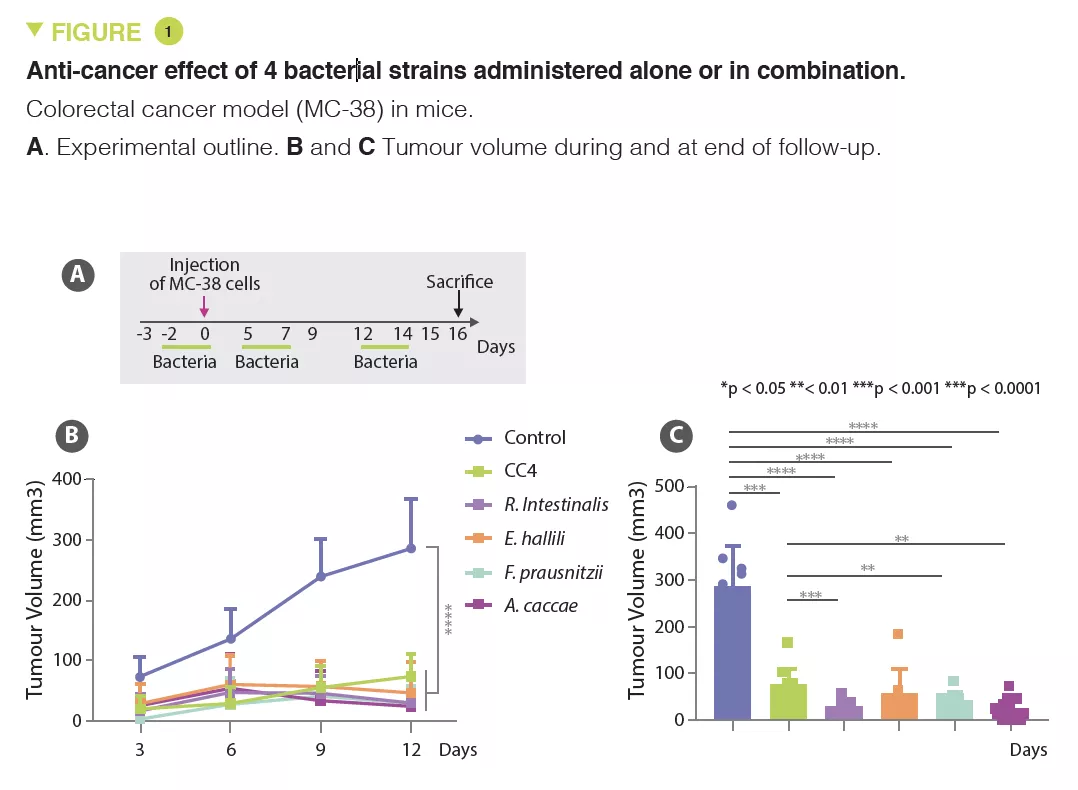
Despite overall success, T-cell checkpoint inhibitors for cancer treatment (anticancer immunotherapy) are still only efficient in a minority of patients. Gut microbiota was recently found to critically modulate anti-cancer immunity and therapeutic response. In this article the authors identified Clostridiales members of the gut microbiota associated with a lower tumour burden in colorectal cancer (CRC) mouse models. Interestingly, these commensal species are also significantly reduced in CRC patients compared with healthy cont rols. Gastric gavage administration of a combination of four Clostridiales strains (CC4) in mice prevented and even successfully treated CRC as stand-alone therapy. This effect was dependent on intratumoral infiltration and activation of CD8+ T cells. Administration of a single strain of Roseburia intestinalis or Anaerostipes caccae was even more effective than CC4. In a direct comparison, the CC4 combination supplementation outperformed anti-PD-1 therapy in CRC and melanoma mouse models. These findings provide a strong preclinical basis for exploring gut bacteria as novel stand-alone therapy against solid tumours.
Gut microbiota plays a major role in the development of the immune system and maintenance of immune homoeostasis. Gut microbiota influence the immune system both on a local and systemic level, maintaining a balanced immune response. In common with a wide range of diseases such as obesity, chronic inflammatory bowel disease, neuropsychiatric disorders, and colorectal cancer (CRC), disruption to the microbial balance (dysbiosis) is implicated.
CRC, as one of the most frequently diagnosed malignant diseases, remains the leading cause of cancer deaths worldwide related to lifestyle (diet, tobacco, alcohol) or obesity risk factors, which are all closely related to changes in the composition of healthy gut microbiota.
Gut microbiota is altered in CRC patients, with an increase in taxa such as Bacteroides or Fusobacterium [2] and a decrease in bacterial Clostridiales taxa compared to healthy individuals [3]. A majority of colorectal tumours have low immunogenicity and therefore do not respond to current T-cell activation immunotherapies. Anti-PD-1, anti-PD-L1 and/or anti-CTLA4 monoclonal antibodies are only effective in 4-5% of CRC tumours with a mismatch repair defect or high microsatellite instability [4]. Recent studies have shown how some members of the gut microbiota are able to modulate the efficacy of anti-cancer treatments [5, 6]. In this study, the authors evaluated whether a cocktail of four bacterial strains, specifically associated with a low tumour burden in an experimental CRC model, could trigger an effective anti-tumour immune response.
Based on animal models, the authors observed that mice with a microbiota low in Clostridiales bacteria (Ruminococcaceae and Lachnospiraceae families) had increased susceptibility to CRC. Based on these analyses, the authors selected a combination of four Clostridiales species (CC4), i.e. Roseburia intestinalis, Eubacterium hallii (Anaerobutyricum hallii), Faecalibacterium prausnitzii and Anaerostipes caccae, whose abundance is decreased in CRC patients, to conduct additional studies on mouse models. CC4 administration had an effect on the mice microbiota, with an increase in species belonging to the Ruminococcaceae and Lachnospiraceae families.
In several solid cancer models, including for CRC, lung and breast cancer, CC4 administration slowed down tumour growth. This benefit was primarily lymphocyte-mediated as it disappeared in mice without mature lymphocytes (Rag2 KO mice). The main candidates are interferon-gamma producing cytotoxic CD8+ T cells, which massively infiltrate the tumour in animals receiving CC4. The protective effect was also observed individually with each of the bacterial strains tested in a CRC model, but with varying degrees of efficacy (Figure 1). The therapeutic effect was unrelated to butyrate producing-bacteria.
Finally, in the CRC model (MC-38), the CC4 cocktail had a superior effect to anti- PD-1 immunotherapy. Moreover, no additive effect was observed among mice having received immunotherapy and the four bacteria.
This study has shown that in addition to having an adjuvant role in the immunotherapy treatment of cancer, some microbiota bacteria exert their own anti-cancer effect in solid tumour mouse models. These results pave the way to the development of cancer treatments in man based on gut microbiota. These therapies could be used alone or in combination with other “conventional” anti-cancer treatments. However, the patient groups that would benefit most from this type of therapeutic approach are yet to be determined.
This study in mice showed that bacterial Clostridiales strains, which are significantly lower in colorectal cancer patients, are effective in stimulating an anti-cancer response to solid tumours. The mechanisms involve activation of CD8+ T cells and are independent of anti-PD-1 immunotherapy.
Overview
By Dr. Vincent Trebossen
Child and Adolescent Psychiatry Department, Robert-Debré Hospital, APHP, Paris, France
By Dr. Pierre Ellul
Child and Adolescent Psychiatry Department, Robert-Debré Hospital, APHP, Immunology-Immunopathology-Immunotherapy, Pitié Salpêtrière Hospital, U959 Inserm, Paris, France
By Dr. Alexis Mosca
Pediatric Gastroenterology Department, Robert-Debré Hospital, APHP, Paris, France
By Pr. Richard Delorme
Child and Adolescent Psychiatry Department, Robert-Debré Hospital, APHP, Human Genetics and Cognitive Functions, Institut Pasteur, UMR3571 CNRS, Paris-City University, Paris, France

Autism Spectrum Disorders (ASD) are complex neurodevelopmental disorders affecting 1% of the general population and characterized by a deficit in social communication and repetitive/stereotyped behaviors. The pathophysiological mechanisms behind ASD are still poorly understood [1]. Thirty to 50% of individuals with ASD present gastrointestinal (GI) symptoms such as abdominal pain, diarrhea, constipation, which affect their quality of life and their global functioning. Interestingly, the occurrence and the severity of GI symptoms are strongly correlated with autistic symptoms [2]. Whereas the etiology of GI symptoms are still unknown, several studies suggested that ASD could result from an imbalance in the gut microbiota (GM) composition [3]. Consistent findings suggest robust interactions between GM and the central nervous system (CNS). GM directly affects neurodevelopment by impacting neurogenesis, neuron survival, brain growth and myelination. The modulation of GM using pro- or prebiotics or fecal microbiota transplantation (FMT) in individuals with ASD shows beneficial and long-term effects on GI symptoms and core autistic symptoms. Larger double-blind randomized trials are however needed to confirm the efficacy of microbial-based therapies in ASD, specifically at an early and critical stage of neurodevelopment
The human gut microbiota (GM) consists of approximately 1013 microorganisms, mainly bacteria, fungi and viruses. GM plays a central role in human health, ensuring intestinal barrier function, modulation of the immune response and metabolic synthesis, but also a direct protection against infections. Dysbiosis characterized by an unbalanced GM has been associated with several diseases such as inflammatory bowel diseases, cancers, diabetes or obesity [4]. Consistent findings also suggest robust interactions between GM and the central nervous system (CNS) [5] (Figure 1).
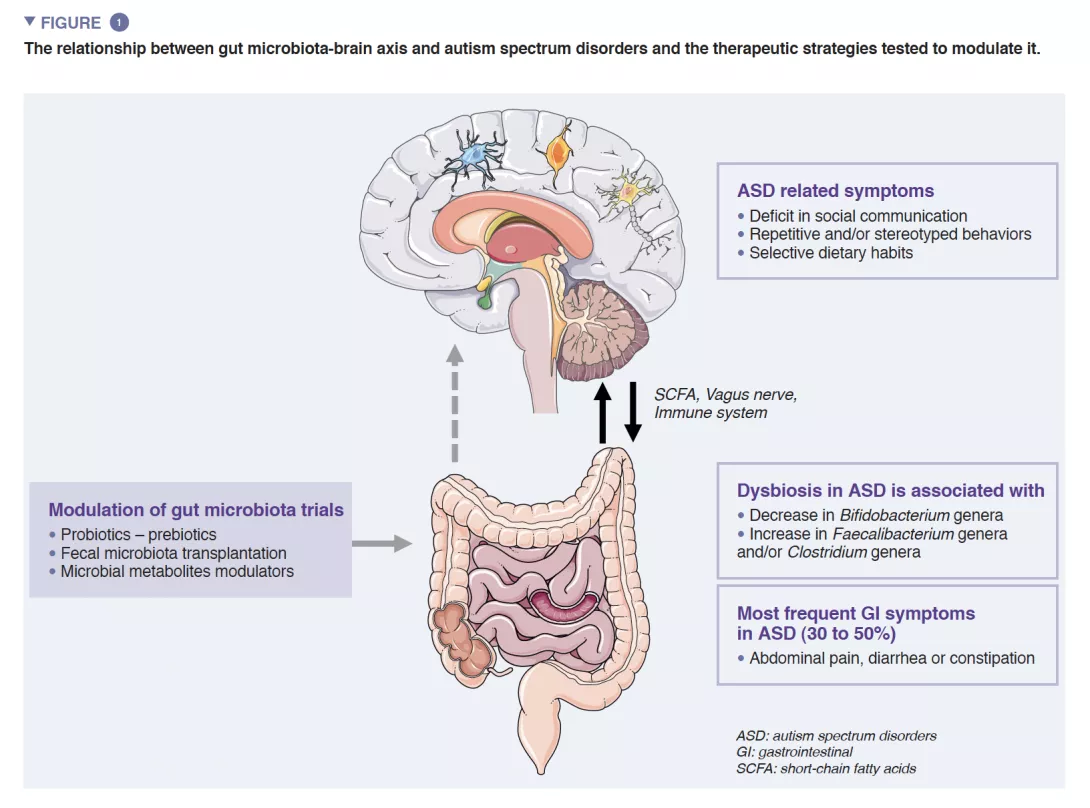
A reciprocal crosstalk between brain and GM is mediated by microbial metabolites (mainly short chain fatty acids) and immune modulators directly by crossing both the blood-gut and the blood-brain barriers and indirectly via the stimulation of the vagus nerve [6]. Evidence supports the involvement of GM in the regulation of both human behaviors and cognitions - specifically socio-communication skills – even if its exact mechanisms are still unknown [5].
ASD are neurodevelopmental disorders characterized by impairments in social communication, social interactions, and repetitive/stereotyped behaviors, of childhood onset affecting approximately 1% of the general population. The determinism of ASD is mainly driven by genetic factors, with a heritability estimated reaching 0.8- 0.9, but pre- and post-natal environmental events may act as precipitating factors or modulators of the symptom severity. The trajectory of brain development at an early stage of life overlaps with those of the GM. This latter begins to develop early after birth and its composition stabilizes into an adultlike profile around the age of 3 years old. GM early composition is deeply influenced by environmental factors such as the place of birth, delivery mode, breastfeeding and xenobiotics (e.g. antibiotics use).
Germ-free mice are a model lacking all microorganisms e.g are microbiologically axenic (no living organisms can be cultured from germ-free mouse specimens). Germfree mouse models are valuable to decipher the mechanisms underlying the roles of GM in neurodevelopment, but also the relationship between microbiome and disease. Studies showed that germ-free mice exhibit i) default in brain-blood barrier permeability; ii) higher brain volume; iii) more immature microglia gene expression and less microglial immune responsiveness; iv) increased myelination ; and v) decreased Brain-Derived Neurotrophic Factor expression and in a subunit of N-Methyl-D-Aspartate receptors [5, 7]. All these data stress the role of GM in blood-brain barrier formation and integrity, neurogenesis, microglia homeostasis, myelination and brain growth/ function.
Germ-free mice displayed autistic-like behaviors such as social avoidance, repetitive/ stereotyped behaviors, lack of interest in social novelty. Some of these behaviors disappeared after colonization by a GM from wild type mice whereas colonization by GM from ASD mouse models increased these behaviors. GM seems indeed crucial for the programming and presentation of social skills and adaptive behaviors [8].
A growing number of evidence shows that GI symptoms are overrepresented in ASD children. GI symptoms such as abdominal pain, constipation, diarrhea are reported in nearly 30-50% of patients with ASD and profoundly impact the quality of life of children [1]. GI symptom severity was correlated with the severity of autistic symptoms and gut dysbiosis is well documented even if there is still no specific signature related to autistic symptoms. Studies exploring GM reported differences in microbiota diversity, and abnormal metabolite patterns when compared to healthy controls. Two recent meta-analyses exploring GM composition in ASD patients reported a decrease in Bifidobacterium and increase in Faecalibacterium and Clostridium genera in ASD patients [9, 10] compared to controls. The exploration of the fecal metabolome also displayed an increase in p-cresol, a bacterial metabolite derived from tyrosine, in individuals with ASD. All together, these data may indicate the potential association between GM abnormalities and GI symptoms in ASD patients.
However, most studies have heterogeneous results and methodological limitations. Merely confounding factors such as different countries with different lifestyles and dietary habits are major drawbacks of these studies. Indeed, a recent study in a large cohort of 247 subjects with ASD did not report direct links between ASD diagnosis or autistic symptoms, and GM dysbiosis. Dysbiosis was associated with a less-diversified diet which is common in patients with ASD [8].
Almost 30-50% of patients with ASD reported GI symptoms such as abdominal pain, diarrhea, constipation. GM dysbiosis in ASD patients have been well explored and confirmed through meta-analysis even if there is no microbiome specific signature [9, 10]. The link between GM dysbiosis and ASD is still poorly understood. Some studies suggest direct links through the GM-brain axis influencing autistic symptoms and GI symptoms. More recently a study suggests that GM dysbiosis in ASD patients is mostly linked with the restrictive diet which is frequent in ASD patients [8].
A growing number of studies explored the potential impact of microbiota-based therapeutic strategies to improve GI symptoms and core symptoms in individuals with ASD.
Probiotics, live microorganisms, have been used in ASD and could have a beneficial effect on patients with ASD. Some preclinical studies showed increased social interactions with probiotics supplementation (Bacteroides fragilis, Lactobacillus reuteri) in mouse models of ASD. The improvement of social communication was linked with an increased oxytocin expression in CNS. In humans, several studies reported positive effects of probiotic treatments on GM composition and GI symptoms in ASD [11]. However, few of them reported an improvement of core autistic symptoms. Most clinical trials providing probiotics in autistic individuals showed inconsistency in terms of probiotics, dosage administration per day or in total, and duration of the whole treatment. Even if some studies suggest that probiotics could be interesting to prevent GI symptoms in ASD patients, the results request replication to guarantee the positive effect of such strategy.
Similarly, the efficacy of prebiotics, such as galacto-oligosaccharide (GOS) or fructo-oligosaccharide have been explored in ASD [12]. Chronically stressed mice showed alteration in GM and a decrease in social interest. Using this mouse model, the administration of prebiotics was associated with increased social interactions in these mice. In humans, the use of GOS associated with a casein-free and gluten-free diet showed improvement in GI symptoms and social interactions together with an increase in GM Bifidobacterium abundance. Appropriate double-blind randomized clinical studies are needed to confirm preliminary evidence.
Fecal microbiota transplantation (FMT) has also been studied in ASD. FMT involves transplanting GM from a donor to modify the GM of the receiver. Its efficacy on Clostridioides difficile infection is now well demonstrated, even in children. A recent exploratory unblinded and non-randomized clinical trial involving 18 children diagnosed with ASD and GI evaluated the effect of microbiota transfer therapy (MTT) - a modified FMT protocol [13]. MTT consisted in a two-week antibiotic treatment, a bowel cleansing, before receiving the MTT treatment which consisted of a high dose through oral or rectal administration followed by an oral maintenance dose for 7-8 weeks. Adverse events at the initiation of vancomycin treatment were observed (disruptive behaviors, hyperkinesia) but disappeared spontaneously after 3 days of treatment. The MTT protocol led to a significant improvement in GI symptoms after the following survey of 8 weeks. More surprisingly, an improvement on core autistic symptoms (stereotyped and repetitive behaviors, social communication skills) had also been observed 8 weeks after MTT. Interestingly, improvement on GI symptoms and autistic symptoms persisted 2 years after treatment and was correlated with GM increased diversity [14]. Two years after MTT, the average reduction of the Gastrointestinal Symptom Rating Scale (GSRS) total score was still over 50%. Changes in autistic symptoms measured with the Childhood Autism Rating Scale - CARS, the Social Responsiveness Scale - SRS, or the Autistic Behavior Checklist - ABC were all positively correlated with percent changes in GSRS scores. These results are not yet confirmed by placebo-controlled doubleblind randomized studies.
Recently, a pilot open label clinical trial in ASD has explored the effect of an oral GI-restricted adsorbent (AB-2004) modulating several GM metabolites. The authors reported a decrease in anxiety-like behaviors in mice, driven by a gut microbial metabolite decrease [15]. The study also presented preliminary results from a clinical trial in which an AB-2004 weight-adjusted dose was administered, for 8 weeks, to 30 adolescents with ASD. At week 8, reduced levels of GM metabolites in plasma and urine were observed. More interestingly, after treatment, less subjects displayed GI symptoms but also ASD-associated behaviors, anxiety, and irritability. There was also a remnant effect with a persistence of the efficacy at 4 weeks after treatment discontinuation [15]. The factors linking clinical improvements and administration of AB-2004 remain to be determined, some indirect factors have not been studied such as the effect of AB-2004 on nutrition changes, immune status or GI function. Larger, double-blind placebo- controlled studies trials are warranted to further dissect the role of AB-2004 in social communication in humans.
In the context of the lack of specific treatment for GI symptoms and autistic symptoms in ASD patients, new well tolerated therapeutic strategies targeting GM or microbial metabolites such as FMT/MTT need to be more performed specifically in early and critical stages of brain development during childhood.
FMT is a new efficient way to modulate GM used in gastrointestinal and endocrine diseases. Fecal microbiota transplantation (FMT) could represent a new and efficient opportunity to modulate/reset GM in ASD. Using mouse models of ASD, FMT efficiently improved autistic like core symptoms such as social communication deficits and stereotyped behaviors. In humans, a pilot open-label trial in adults with ASD suggested that FMT could improve not only GI symptoms but also social reciprocity and stereotyped behaviors. There was a remnant effect with a persistent efficacy 2 years after treatment discontinuation. Interestingly the efficacy was associated with an increased GM diversity. FMT could be a new cutting-edge therapeutic strategy to treat GI and behavioral symptoms in ASD children, and merely open new avenues toward physiology of social communication in humans.

The impact of gut microbiota on the neurodevelopmental trajectory of children remains largely unexplored. At this point, there are no studies exploring the impact of gut microbiota modulation on the early phase of child development, particularly on social communication skills. Preliminary studies in adults and children have shown that GM modulation leads to improvement in GI symptoms and autistic behaviors. Based on these data, we will conduct a double-blind clinical trial in younger children with ASD at a very early stage of their cognitive development (before age 6). We aim to understand whether FMT performed at a critical developmental period will result in significant longterm improvement in the child’s developmental trajectory.

