From diarrhea to chronic diseases: the well-documented consequences of antibiotic-related gut microbiota dysbiosis
Antibiotic treatment may sometimes take place without any obvious short-term side effects. Nevertheless, the dysbiosis triggers diarrhea for up to 35% of patients; in the long term, antibiotic-induced microbiota alterations may represent a risk factor for allergic, autoimmune or metabolic diseases.
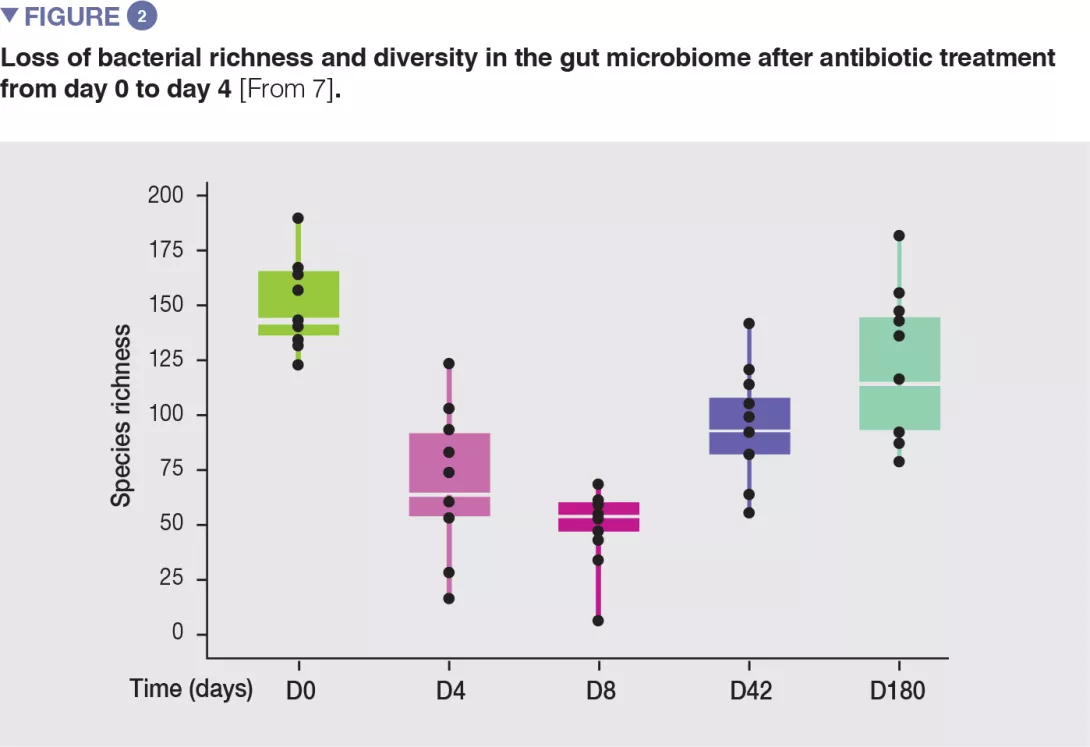
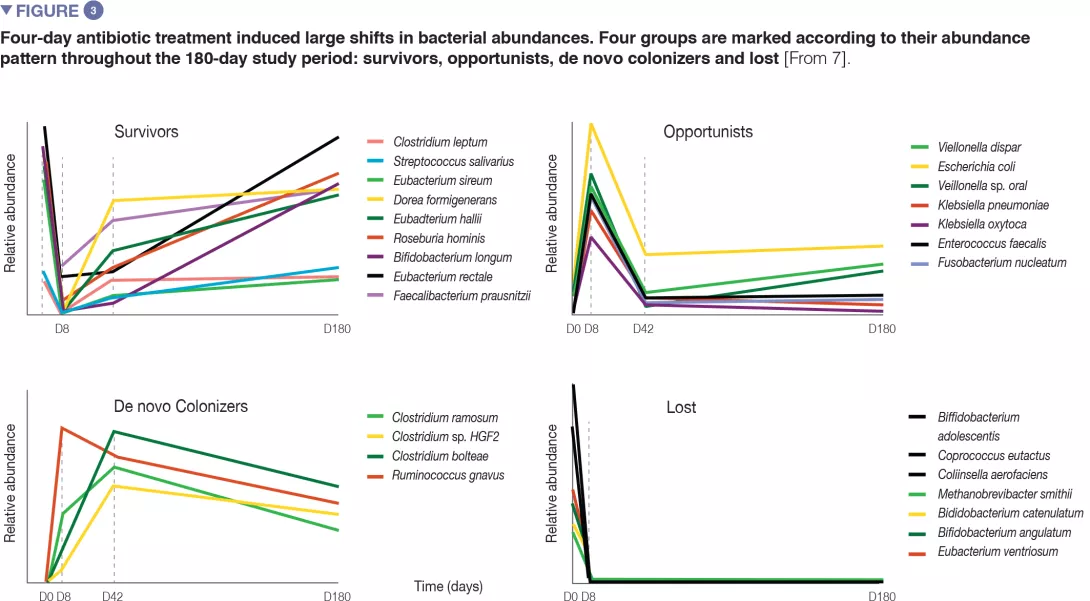
Antibiotics are a powerful tool in the fight against bacterial infections. However, research has also documented detrimental effects on the trillions of commensal bacteria that live in the intestinal tract. This resultant dysbiosis renders the gut microbiota less able to fulfil its protective functions. In the short term, dysbiosis leaves the door open for opportunistic pathogens and the selection of multi-resistant bacteria. In the long term, the gut microbiota, despite having a certain degree of resilience, can sometimes fail to fully restore itself;1,2 this is understood to pave the wave to a range of diseases. Recent research has shown that antibiotics may alter the bacterial diversity and abundance of the normal microbiome and that this impact may be prolonged (typically 8-12 weeks after antibiotics have been discontinued).3,4
35%
Diarrhea occurs in up to 35% of patients who receive antibiotics3, 5, 6
Diarrhea, the most common adverse effect of antibiotics
As the main short-term consequence, some patients treated with antibiotics experience a change in their intestinal transit, most often resulting in diarrhea. The incidence of antibiotic-associated diarrhea (AAD) depends on several factors (age, setting, type of antibiotic, etc.) and may range between 5 and 35% of patients taking antibiotics.3,5,6

Expert opinion
Antibiotics disrupt the protective intestinal microbiota, which can lead to unintended consequences including antibiotic-associated diarrhea (in up to 35% of patients) and the development of antibiotic resistant strains of pathogens that are of global concern in regards to increased healthcare costs and mortality.
Among children this percentage can reach up to 80%.3 Most of the time, the diarrhea is purely functional, caused by the antibioticinduced dysbiosis. It is usually of mild intensity and is self-limiting, lasting 1-5 days. Antibiotics displaying a broader spectrum of antimicrobial activity like clindamycin, cephalosporins, and ampicillin/amoxicillin are associated with higher rates of diarrhea.6
Antibiotics are an extraordinary scientific discovery that saves millions of lives but their excessive and inappropriate use has now raised serious concerns for health, notably with antibiotic resistance and microbiota dysbiosis. Let’s take a look at this dedicated page:
The ambivalent role of antibiotics
The particular case of C. difficile
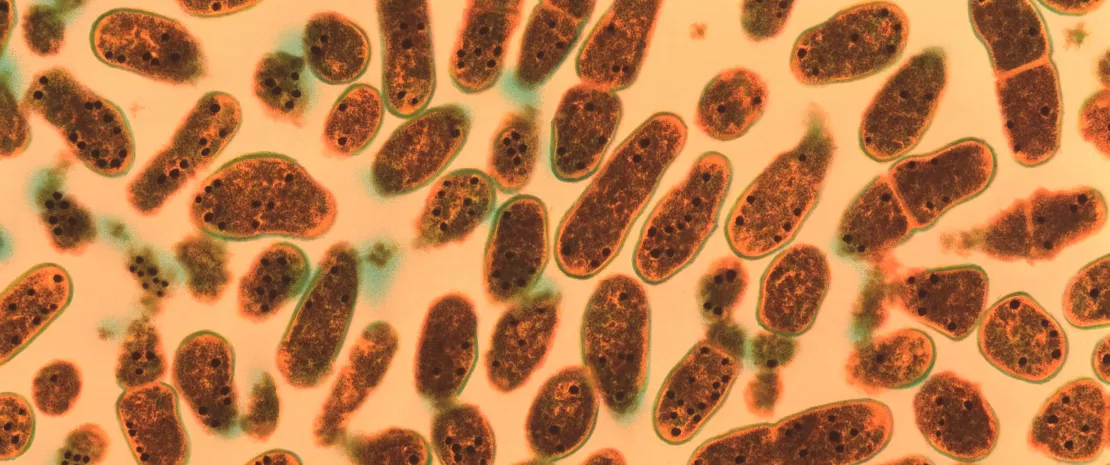
In 10 to 20% of cases, diarrhea results from infection with Clostridioides difficile (formerly known as Clostridium difficile) colonizing the microbiota.6 This bacterium, which persists in the environment via spores, is a gram-positive, spore-forming, obligate anaerobe. Infection occurs via spores ingestion. Under specific circumstances (e.g., antibiotic-induced dysbiosis), the spores may germinate and vegetative bacterial cells of this opportunistic pathogen may colonize the intestines. In the infective phase, C. difficile produces 2 toxins that damage the colonocytes and trigger an inflammatory response with a variety of clinical outlooks, ranging from moderate diarrhea to pseudomembranous colitis, toxic megacolon and/or death.
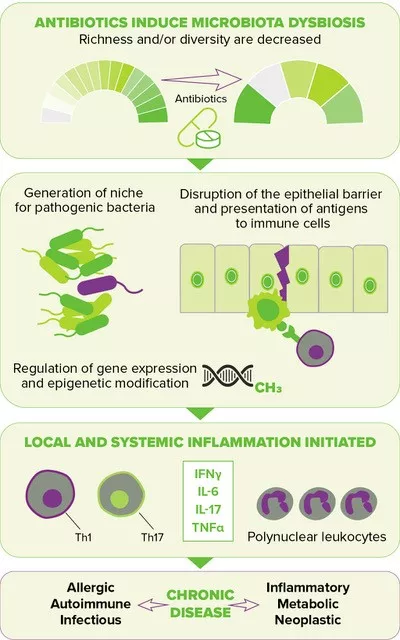
FIGURE 2. Downstream effects of antibiotic-induced gut dysbiosis. (source : adapted from Queen et al., 202010)
1/3
Nearly 1/3 of AAD cases are due to C. difficile3
Most recognized common risk factors for C. difficile infection (CDI) include age > 65 years, use of proton pump inhibitors, comorbidities and of course antibiotic use. The latest is the most relevant modifiable risk factor for CDI. The association of antibiotics with CDI has been established in hospitals and more recently in community settings,7 where the risk of infection varies from intermediate for people exposed to penicillins, high for these exposed to fluoroquinolones and highest for those receiving clindamycin. As for tetracyclines, they trigger no increased risk.8 In a hospital setting, the highest risk of developing CDI was observed for cephalosporins (from 2nd to 4th generations), clindamycin, carbapenems, trimethoprim sulfonamide, fluoroquinolones and penicillin combinations.9
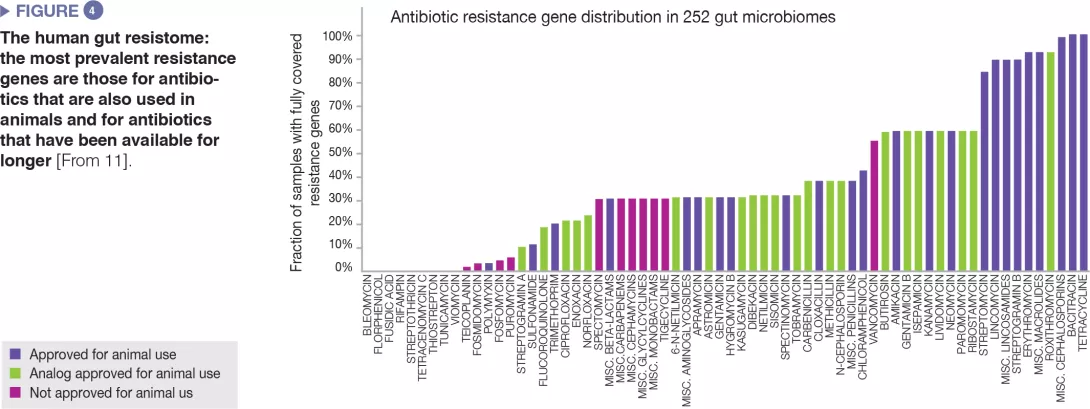
When the gut microbiota becomes a reservoir of antibiotic resistance
When exposed to antibiotics, microbial communities respond in the short term not only by changing their composition, but also by evolving, optimizing and disseminating antibiotic resistant genes. The human gut microbiota overly exposed to antibiotics is now considered a significant reservoir of resistance genes, in adults as well as in children.2 By contributing to the growing difficulty to combat bacterial infections, antibiotic resistance has become a major public health concern.
An open door to non-communicable diseases
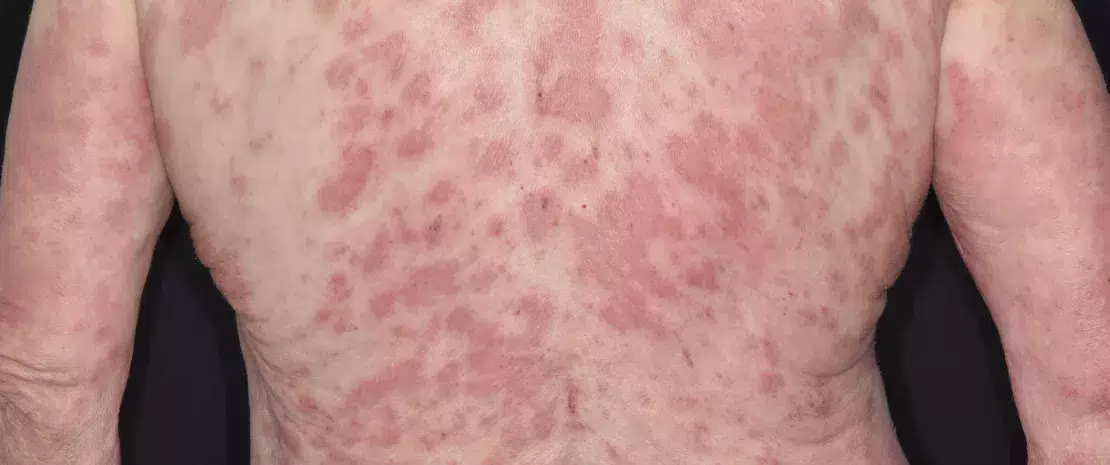
Disruption of the gut microbiota resulting from antibiotic exposure is also suspected of increasing the risk of several chronic diseases by elevating inflammatory responses locally and systemically, thereby leading to a deregulated metabolism and compromised immune homeostasis10 (Figure 2, page 4). The perinatal period, characterized by the development of the immune system along with the maturation of the gut microbiota, has been shown to be a particularly sensitive time, one during which antibiotic-driven dysbiosis translates into long-lasting health effects, i.e. a higher risk of diseases later in life, including inflammatory bowel diseases (e.g., Crohn’s disease), atopic diseases (e.g., asthma) and metabolic disorders (e.g., type 2 diabetes, obesity).
Clinical case
by Lynne V. McFarland, PhD
-
53-year old woman consulted her physician with a 3-day history of respiratory tract symptoms (cough, sore throat and runny nose) with fever and fatigue. No co-morbidities and was otherwise healthy. Her physician prescribed a sputum sample and a 10-day course of oral cefaclor (500 mg, b.i.d). The sputum cultures came back negative for pathogens.
-
She was admitted at hospital on the 3rd day of the antibiotics because she developed acute diarrhea (with six watery stools per day and abdominal cramping) and unresolved respiratory symptoms. Laboratory cultures (sputum and stool) were negative for pathogens. She was asked to discontinue her antibiotics, but the diarrhea continued for the next two days.
-
Her physician prescribed erythromycin (500 mg, t.i.d.) and a probiotic for one week. Her respiratory symptoms and diarrhea resolved within four days and she was discharged one day later with no complications.
Each year, since 2015, the WHO organizes the World AMR Awareness Week (WAAW), which aims to increase awareness of global antimicrobial resistance.
Held on 18-24 November, this campaign encourages the general public, healthcare professionals and decision-makers to use antimicrobials carefully, to prevent the further emergence of antimicrobial resistance.
 Microbiota at the forefront of antibiotic resistance
Microbiota at the forefront of antibiotic resistance