Skin microbiota #10
By Pr. Markku Voutilainen
Turku University Faculty of Medicine; Turku University Hospital, Department of Gastroenterology, Turku, Finland


By Pr. Markku Voutilainen
Turku University Faculty of Medicine; Turku University Hospital, Department of Gastroenterology, Turku, Finland

The meibum prevents the evaporation of the eye’s tear film, allows homeostasis of the ocular surface and has its own microbiota. Does this microbiota, and those of ocular surfaces, change with ageing? This study shows that eyelid-skin samples from young subjects had low α-diversity (Shannon index) and that Probionibacterium acnes and Staphylococcus epidermidis were the dominant species. Meibum and conjunctival-sac microbiota were different from that of the skin and characterized by a high α-diversity index consisting in a large number of bacterial species. In elderly subjects, Corynebacterium sp. and Neisseriaceae were the predominant taxa on eyelid skin and α-diversity Shannon index was significantly reduced at their meibum and conjunctival sac. Authors conclude that meibum’s microbiome is indeed altered with ageing, equally in men and in women.
We know that gut microbiome changes with age, as well as oral and skin ones. Which of them is best to predict ageing? The authors evaluated the microbiome diversity from almost 9 000 skin, saliva and intestinal samples from healthy people across 10 studies. Taxa enriched in young individuals (18 to 30 years) tended to be more abundant and prevalent than taxa enriched in elderly persons (> 60 years); and ageing may be linked to a loss of key taxa. Compared with gut and oral microbiome, the skin one was the best predictor of age (mean 3.8 yrs ± 0.45 SD). Authors identified genera and families including anaerobic bacteria (Mycoplasma, Enterobacteriaceae, Pasteurellaceae) that negatively correlated with age. Age-related changes in skin physiology (decreased sebum production, increased dryness) and host immune reactions may cause these microbiome changes.
By Pr. Markku Voutilainen
Turku University Faculty of Medicine; Turku University Hospital, Department of Gastroenterology, Turku, Finland
Could gut and vaginal microbiomes predict the stage of endometriosis? According to this study, no differences were detected between samples from patients and healthy control subjects for follicular and menstrual phases of the menstrual cycle. At an individual level, the distribution of community state types differed significantly between the two menstrual phases. Among patients, Operational Taxonomic Units (OTU) of genus Anaerococcus (notably A. lactolyticus and A. degenerii) significantly differed between disease stages 1-2 and stages 3-4 (rASRN - revised American Society for Reproductive Medicine - classification). Authors conclude that vaginal microbiota may predict the stage of the disease
Vaginal dysbiosis may be linked to the development, progression and stability of cervical cancer (CC), but it is unclear whether the relationship between the two is causal or correlative. To find out if cervicovaginal bacteria also have an anticancer effect, the use of microorganisms in the treatment of cancer (especially CC) was discussed in this article: probiotics, bacteria-based immunotherapy, bacterial toxins and spores, vectors in gene therapy, and inhibitors of tumor angiogenesis. Some bacteria inhibit CC by activating NK cells and dendritic cell maturation. Other mechanisms are production of cytotoxic compounds, regulating immune cell differentiation, and inhibiting cancer cell migration. In conclusion, genetically engineered bacteria may be an effective treatment for CC in the future. Larger sample size studies are needed to assess this option.
Congress review
By Pr. Kristin Verbeke
Laboratory for Digestion and Absorption, Translational Research Center for Gastrointestinal Disorders (TARGID), Department of Clinical and Experimental Medicine, Leuven, Belgium

This year’s Gut Microbiota For Health World Summit was held in Madrid under rather exceptional conditions due to the rising Covid-19 epidemic. As many participants could not physically attend the meeting, this report is somewhat more extensive than the standard congress reviews in this Newsletter.
The 9th edition of this symposium addressed various aspects of the microbiota including dietary and non-dietary factors shaping the gut microbiota and the role of microbiota on brain function and in modulating the immune system. In an introductory keynote lecture, Colin Hill (Cork, Ireland) underlined some roadblocks in the clinical translation of microbiome research. He made an appeal to adopt consensus definitions and to use precise language. The goal is to make microbiome science more numerate and use actual numbers of bacteria rather than only relative abundances and proportions. It tends to address the complexity related to microbiome individuality and to monitor the complexity of figures in order not to overwhelm readers.
Furthermore, he pointed out that the choice of methodology, such as biological vs. in silico methods, significantly affects the results.
For example, the faecal microbiome is only a very blurred approximation of the intestinal microbiome, intestinal transit times affect the composition of the microbiome and the conversion of in silico to in vivo methods is not always straightforward. Microbiome research should be a science where the highest standards are applied rather than a belief system.
Previous cross-sectional studies already indicated that the composition of the faecal microbiota depends on dietary patterns. In particular, people that consume a plantbased diet have a more diverse microbiota with a higher proportion of short-chain fatty acids (SCFA) producing bacteria compared to people on a western-type diet high in refined carbohydrates and fat. Changing diet from a standard American diet to a plant-based diet modified the microbiota and markedly improved the metabolic outcome of obese subjects as reported by Hana Kahleova (Washington DC, USA). Participants that switched to a plant-based diet for 16 weeks lost 5.8 kg of body weight, of which about two thirds was fat, and had improved insulin sensitivity compared to the control group that did not adapt its dietary pattern. Faecal Bacteroidetes and Faecalibacterium prausnitzii increased on the vegan diet, while Bacteroides fragilis decreased on both diets but less on the vegan diet. Furthermore, changes in bacterial composition correlated to changes in metabolic parameters.

As highlighted by Rinse K. Weersma (Groningen, The Netherlands), drugs interact with the intestinal microbiota according to different scenarios. Some drugs like proton pump inhibitors (PPI) affect the microbiota composition and functionality. The higher pH locally in the gut due to PPI intake results in “oralisation” of the gut microbiota as oral bacteria manage to penetrate deeper in the gastrointestinal tract. The oral antidiabetic metformin also affects the gut microbiota composition by increased numbers of Akkermansia muciniphila and SCFA production which contributes to its antihyperglycemic effect. Immune therapeutics do not directly affect the microbiome but as the microbiome is involved in immune homeostasis, it indirectly determines the response to those anticancer drugs. Furthermore, the microbiota also modifies the activity of drugs by activating or inactivating them or by influencing their toxicity. For example, the conversion of levodopa by the gut microbiota makes it less bioavailable to the brain which may explain part of the variable responses of patients to this drug. To be active, the prodrug sulfasalazine needs to be split by bacterial azo-reduction in the colon into 5-ASA and sulphapyridine whereas the cardiac glycoside digoxin is inactivated by microbial metabolism. Finally, the gut microbial conversion of the oral antiviral drug brivudine into bromovinyluracil is involved in its toxicity.
Athanasios Typas (Heidelberg, Germany) highlighted that the impact of non-antibiotic drugs on the microbiota is extensive. By screening 1,200 marketed drugs in vitro against 40 representative gut bacterial strains, at least a quarter of non-antibiotic drugs with human targets was found to inhibit at least one strain.[1] This in vitro inhibition was reflected in the side effects of the drugs in humans and was concordant with existing clinical trials indicating the relevance of the screening strategy. Remarkably, there was a substantial overlap in susceptibility of gut bacterial strains for human-targeted drugs and antibiotics which was attributed to the fact that the same pumps, transporters and detoxification mechanisms are used for both groups of drugs. These results imply that polypharmacy may be a strong driver for antibiotic resistance.
In the last 30 years, very little progress has been achieved in therapy for mental health. John Cryan (Cork, Ireland) argued that the gut microbiota might provide a new target to improve brain health, despite it still being early days, as the gut microbiota affects brain health during different stages of life. The mode of birth that impacts the intestinal microbiota has been associated with neurodevelopmental disorders.[2] Mice born via caesarean section exhibit an increased stress response, higher anxiety and deficits in sociability. These effects can be reversed by targeting the gut microbiota. The fact that also germ-free mice show inappropriate brain development as evidenced by deficits in fear memory, increases in visceral pain and social deficits, corroborates a role for the gastrointestinal microbiota. Furthermore, during early adolescence, the brain is sensitive to microbial signals. Mice that received a high fat diet during the adolescent period had long-lasting differences in gut microbiota composition in adulthood, together with differences in expression of genes related to neuroinflammation or neurotransmission, although no overt behavioural changes in adulthood were observed.[3] In aged male mice, shifts in the microbiota towards a profile previously associated with inflammatory diseases were associated with increased gut permeability, peripheral inflammation and behavioural changes including deficits in spatial memory and increased anxiety-like behaviour.
Well known as a “happiness” hormone, serotonin actually has a much more complex biological function. It is involved in bone density and in neural, platelet and gastrointestinal function which makes it an attractive intervention point to improve health. The vast majority of serotonin is located in gastrointestinal tissues. Jonathan Lynch (Los Angeles, USA) pointed out that the gut microbiota critically regulates serotonin production by the host. Especially indigenous spore-forming bacteria promote serotonin biosynthesis via production of soluble metabolites that directly signal to colonic cells. This bacterial mediated induction of serotonin regulates gastrointestinal motility and platelet function in mice.[4] Furthermore, luminal intestinal serotonin concentrations also modulate bacterial colonisation in the gut. The relative abundance of spore-forming bacteria, in particular Turicibacter sanguinis, increases when luminal intestinal serotonin levels are elevated. T. sanguinis expresses a receptor homologous to the mammalian serotonin transporter SERT that allows importing of serotonin resulting in expression of sporulation factors and membrane transporters. These effects are reversed by exposure to fluoxetine, a serotonin reuptake inhibitor.
Newborn babies acquire microbes at birth through vertical transmission from their mothers. This postnatal colonisation is assumed to be the main stimulus to the development and maturation of the immune system. Using a model of transiently colonising pregnant female mice, Kathy McCoy (Calgary, Canada) demonstrated that already during pregnancy the maternal gut microbiota shapes the function of the immune system of the offspring. Germ-free pups born to dams that were transiently colonised had increased levels of innate immune cells in the gut and increased expression of genes encoding epithelial antibacterial peptides and metabolism of microbial molecules compared to pups born to germ-free dams.[5] This maternal microbiota-mediated education of the immune system requires maternal antibodies that are transmitted to the offspring during pregnancy and in milk. Furthermore, the maternal gut microbiota protects pups from excessive inflammation. LPS-administration elicited a huge inflammatory response in pups born to germ-free dams whereas this response was blunted in pups born to colonised dams.
The period between birth and weaning, i.e. induction of a more diverse diet, is important for the ontogeny of the immune system, as highlighted by Gérard Eberl (Paris, France). The expansion of the gut microbiota that occurs at weaning induces a strong immune response that is associated with the induction of regulatory T cells [6]. Exposure of germ-free mice to microbes before weaning results in such (normal) immune reaction whereas no reaction occurs when the mice are exposed to microbes only after weaning, indicating that the immune system needs to be exposed to microbes in a specific time window.
Pathological imprinting resulted in increased susceptibility to immune pathologies later in life. How the immune system remembers needs further clarification. Hints in the literature suggest towards epigenetic regulation of immunes genes and imprinting of the expression of genes in myeloid cells or stromal cells. Most likely, many different cells are imprinted, the significance of which needs to be investigated.
Commented articles - Children's section
By Pr. Emmanuel Mas
Gastroenterology and Nutrition Department, Children’s Hospital, Toulouse, France
Commentary on the original article of Lindfors et al. (Gut 2019) [1]
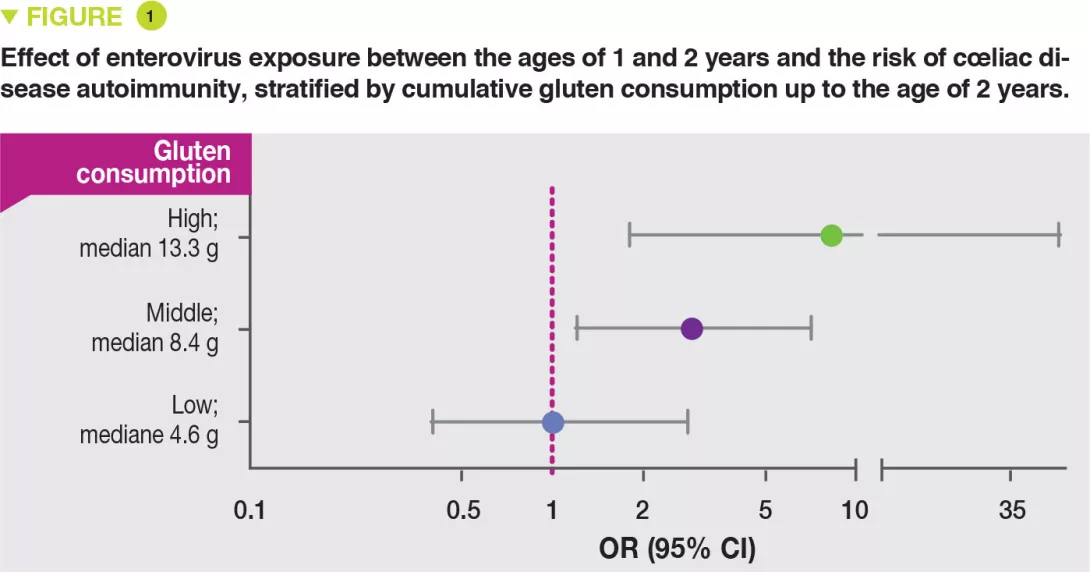
A higher gluten intake, frequent gastrointestinal infections and adenovirus, enterovirus, rotavirus and reovirus have all been proposed as environmental triggers of coeliac disease (CD). However it is not known whether an interaction exists between the quantity of gluten ingested and exposure to viruses in the development of CD. This study sought to determine whether distinct viral exposures, alone or associated with gluten, increased the risk of CD autoimmunity in genetically predisposed children. It was concluded that frequent exposure to enteroviruses between the ages of 1 and 2 years was indeed associated with increased risk of CD autoimmunity, indicating a cumulative effect of the interaction between enteroviruses and higher gluten intake.
Coeliac disease is an autoimmune pathology which occurs in genetically predisposed individuals of genotype HLA DQ2 and/ or DQ8-positive. It is characterised by the presence of villous atrophy and lymphocyte infiltration of the epithelium of the small intestine. Gluten present in the diet induces an autoimmune response directed against tissue transglutaminase. The appearance of anti-transglutaminase antibodies (ATA) indicates the presence of coeliac disease autoimmunity
The rise in the incidence of autoimmune diseases has led to suspect that environmental factors may have a role in their pathogenesis. Observational studies suggest that viral infections could cause a loss of oral tolerance to gluten and the development of coeliac disease.
It is a nested case-control study in the TEDDY (The Environmental Determinants of Diabetes in the Young) birth cohort which included 8,676 children before the age of 4 and a half months followed to the age of 15 years. The principal objective of this cohort was to identify the genetic and environmental factors associated with type 1 diabetes and coeliac disease. After forming pairs matched for family history of type 1 diabetes, sex and study inclusion site, 83 pairs (child with predisposition (case) and control) were retained in the final analysis for whom faecal virome data were available after introduction of gluten. Of these pairs, 16 had a family history of type 1 diabetes. During follow-up, 28 of the coeliac disease autoimmunity cases developed coeliac disease.
Stool samples were collected every month from the age of 3 months to 2 years; tests for enterovirus, adenovirus, astrovirus, norovirus, reovirus and rotavirus were performed. Every 3 months, a food questionnaire was used to collect information on breast-feeding and the age of introduction of gluten-containing foods. A 3-day record of food intake enabled a calculation of the quantities of gluten ingested at 6, 9, 12, 18 and 24 months.
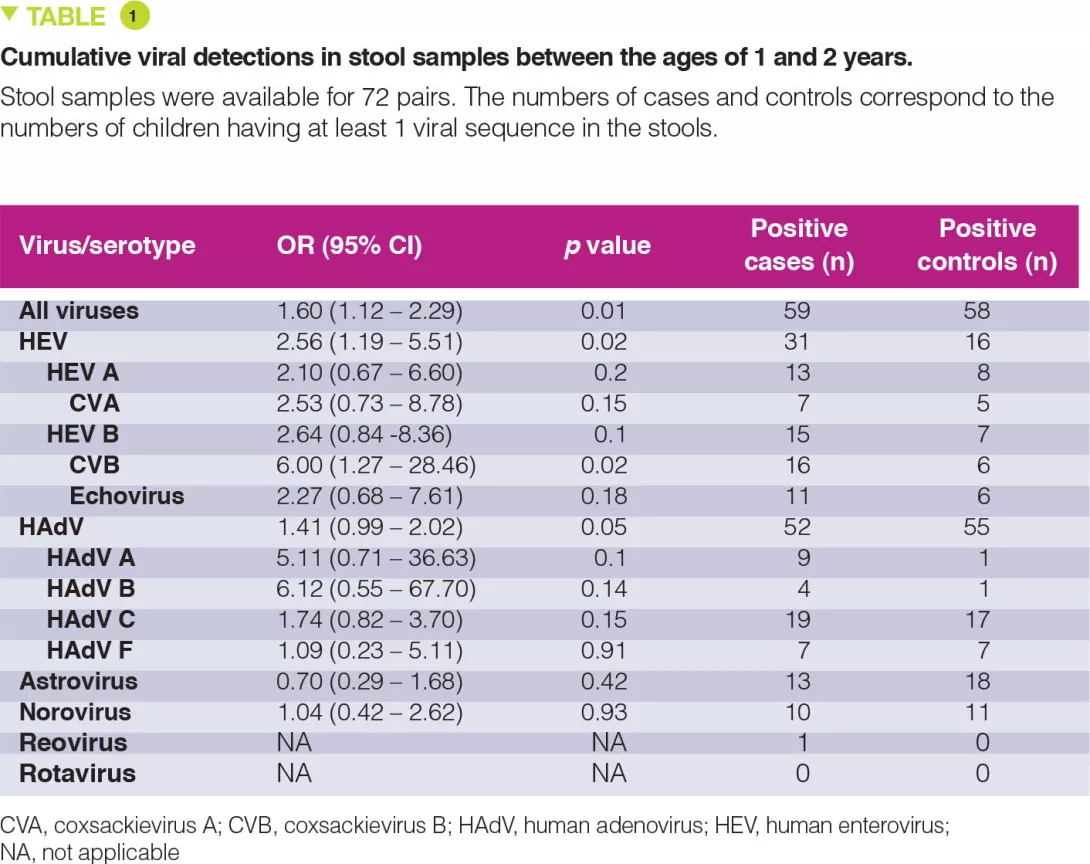
The percentage of stool samples positive for any virus fluctuated from 22 to 50%, without any age-related peak and for the enteroviruses this ranged from 0 to 21% after 6 months. Between 1 and 2 years, enteroviruses were detected in 31 cases versus 16 controls (Table 1). The cumulative number of stool samples positive for any virus was associated with an increased risk for coeliac disease autoimmunity (OR 1.60; p = 0.01), with a stronger association conferred by the enteroviruses (OR 2.56; p = 0.02).
The risk of coeliac disease autoimmunity was not increased by viral infections occurring after the age of gluten introduction while breast-feeding was still continuing. In contrast, after weaning, in stool samples collected between the ages of 1 and 2 years after gluten introduction, both the cumulative number of viruses detected (OR 1.41; p = 0.05) and also the numbers of enteroviruses (OR 2.47; p = 0.03) were associated with the risk of coeliac disease autoimmunity. There was a significant interaction between the presence of enterovirus detected between 1 and 2 years and the quantity of gluten ingested up to the age of 2 years in the risk of coeliac disease autoimmunity (p = 0.03). It is suggested that this increases with the amounts of gluten ingested: high (OR 8.3), middle (OR 2.9) and low (OR 1.0) (Figure 1).
Environmental factors play a role in cœliac disease.
Enterovirus exposure is a risk factor for the development of anti-transglutaminase antibodies in HLA DQ2 and/or DQ8-positive children.
This risk is potentiated by the intake of large amounts of gluten in the diet.
The results of this study indicate the value of preventing the appearance of coeliac disease auto-antibodies in children at risk. This could be done by carefully monitoring the amounts of gluten ingested, in particular in case of exposure to enteroviruses, especially when the child is no longer being breast fed.
This study demonstrated an association between gastro-intestinal exposure to enteroviruses and the risk of coeliac disease autoimmunity in genetically at-risk children. This risk increases as greater amounts of gluten are ingested.
Commented articles - Adult's section
By Pr. Harry Sokol
Gastroenterology and Nutrition Department, Saint-Antoine Hospital, Paris, France

Commentary on the article of Ghosh et al. Gut 2020 [1]
Ageing is accompanied by a deterioration of many bodily functions and inflammation, which collectively contribute to frailty. It has already been shown by the authors and other teams that frailty is associated with cha nges in the gut microbiota, and more especially in the context of a poorly diversified diet. The Mediterranean diet is associated with good health. In this study the authors sought to determine if 12 months of Mediterranean diet, known to be associated with good health, could modify the gut microbiota, reduce frailty and improve cognitive function. The gut microbiota was profiled in non-frail or pre-frail subje cts in five European countries before and after the adoption for 1 year of a Me diterranean diet tailored for elderly subjects (NU-AGE diet). The results showed that it is feasible to improve the usual dietar y regime in order to modulate the gut microbiota, and thus promote healthier ageing.
Frailty which accompanies ageing involves the failure of several physiological systems and constant activation of the innate inflammatory immune response. Frailty can include the development of chronic lowgrade inflammation, impaired cognitive function, sarcopenia and the development of chronic diseases such as diabetes and atherosclerosis. The modification of dietary regimens such as the adoption of a Mediterranean diet has been suggested as a key therapeutic strategy to combat frailty.[2] The Mediterranean diet is characterised by the consumption of larger amounts of vegetables, pulses, fruits, nuts, olive oil, fish and the consumption of smaller amounts of red meat, dairy products and saturated fats. The adhesion to this type of diet is associated with reduced mortality and increased anti-oxidant activity, as well as a reduction in the incidence of several diseases and inflammation.
Several studies have shown that the adoption of this diet is related to a reduction in frailty. Beyond the inverse relationship with disease, closer adhesion to a Mediterranean diet was associated with beneficial changes in the composition of the gut microbiota (reduction in proteobacterial abundance, increased production of short chain fatty acids [SCFAs]). As a general rule however, few elderly subjects follow this type of diet and a large number suffer because of a restricted diet associated with a low-diversity gut microbiota. Changing this is a major challenge, in particular concerning persons in care homes.
In previous studies the authors used bioinformatic analysis to identify specific microbial taxa which are gradually lost in the transition from a high-diversity microbiota of healthy subjects to a low-diversity microbiota of frail subjects. In a recent 6-month dietary intervention study in elderly individuals given supplementation with 5 prebiotics (up to 20 g/day), several microbial taxa were modified, but no change was noted in the overall diversity of the microbiota nor in the inflammatory markers. The authors therefore concluded that a more drastic dietary intervention was necessary. The dietary intervention NU-AGE project aimed to study the effect of administration of a personalised Mediterranean diet for 12 months in a large cohort of over 1,200 persons aged 65 to 79 years, distributed across five European countries. A significant relationship was observed between increased adherence to the Mediterranean diet and global cognitive capacity and improved episodic memory.[3]
Moreover, it was shown that greater adherence reduced the rate of bone loss in individuals with osteoporosis and improved innate immune function, blood pressure and arterial stiffness.[4-6] In the study described here, the authors analysed the gut microbiota of a sub-group of study subjects.
A total of 612 subjects were analysed (289 controls: 145 males, 144 females and 323 on a Mediterranean diet: 141 males, 182 females). At baseline, differences in terms of diet and microbiota were observed between the various countries. Relationships between the Mediterranean diet and the gut microbiota were revealed. Among the taxa associated with good adherence with the Mediterranean diet (DietPositive), we find an over-representation of species such as Faecalibacterium prausnitzii, Eubacterium and Roseburia, a majority of which are associated with good health (including the production of SCFAs and anti-inflammatory effects). Inversely, certain taxa are depleted in case of good adherence to this diet, some of which have been linked to type 2 diabetes, colorectal cancer, cirrhosis or chronic inflammatory bowel disease. Taken together, these results suggest that adherence to a Mediterranean diet can modulate the microbiota in a direction positively associated with health.
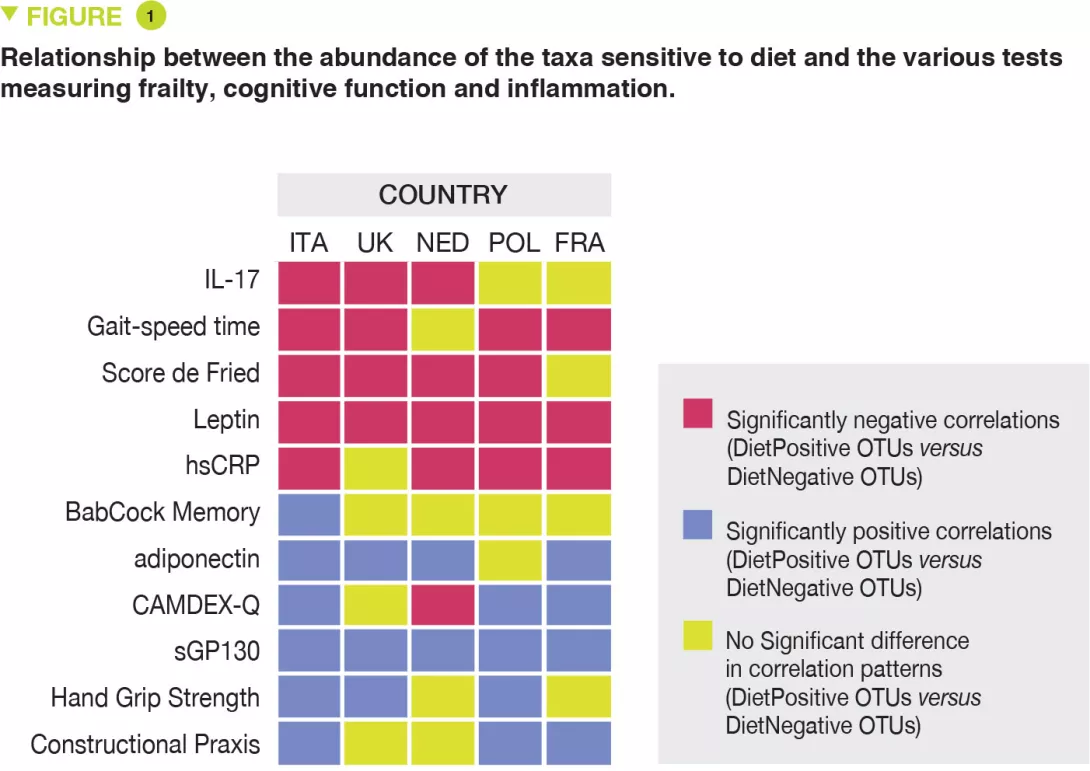
Lastly, the authors observed that the abundance of DietPositive taxa were negatively correlated with some inflammatory markers (high-sensitivity CRP (hsCRP) and IL-17), and with clinical scores associated with increased frailty (Fried scores, gait speed time). In contrast, the abundance of these taxa was positively correlated with the improvement in cognitive function (Constructional Praxis score, Babcock memory score) and reduced frailty (hand grip strength) and two anti-inflammatory markers (adiponectin and sGP130). The opposite trend was observed with DietNegative taxa (Figure 1). Analysis of the inferred microbial metabolite profiles indicated that the diet-modulated change in microbiota was associated with an increase in production of short/branched chain fatty acids and a lower production of secondary bile acids, p-cresols, ethanol and carbon dioxide.
Ageing is associated with a deterioration in numerous functions and inflammation which lead to frailty.
A Mediterranean diet induces an increase in the abundance of bacterial taxa which are associated with a reduction in frailty.
The role of an increase in the production of some bacterial metabolites, including SCFAs, is suggested in the positive effects of the diet.
These results confirm that dietary intervention is an effective means of improving health, at least partially, via a modulation of the gut microbiota. Of course we can recommend that elderly subjects adopt a Mediterranean diet, but the feasibility of this type of dietary intervention is questionable in the long-term. As this study has identified bacteria associated with the beneficial effects of the Mediterranean diet, it lays the groundwork for their use in the form of next-generation probiotics. This type of approach based on bacteria from the gut microbiota should be tested in this indication.
This study highlights the complex interactions between diet, the gut microbiota and health. It suggests that the beneficial effects of a Mediterranean diet on the health of elderly subjects is due, at least in part, to a modulation of the gut microbiota.

"Good Science - worth a read..." -@ethicos2013 (From Biocodex Microbiota Institute on X)
Overview
By Pr. Gianluca Ianiro
Digestive Disease Center, Fondazione Policlinico “A. Gemelli” IRCCS, Rome, Italy
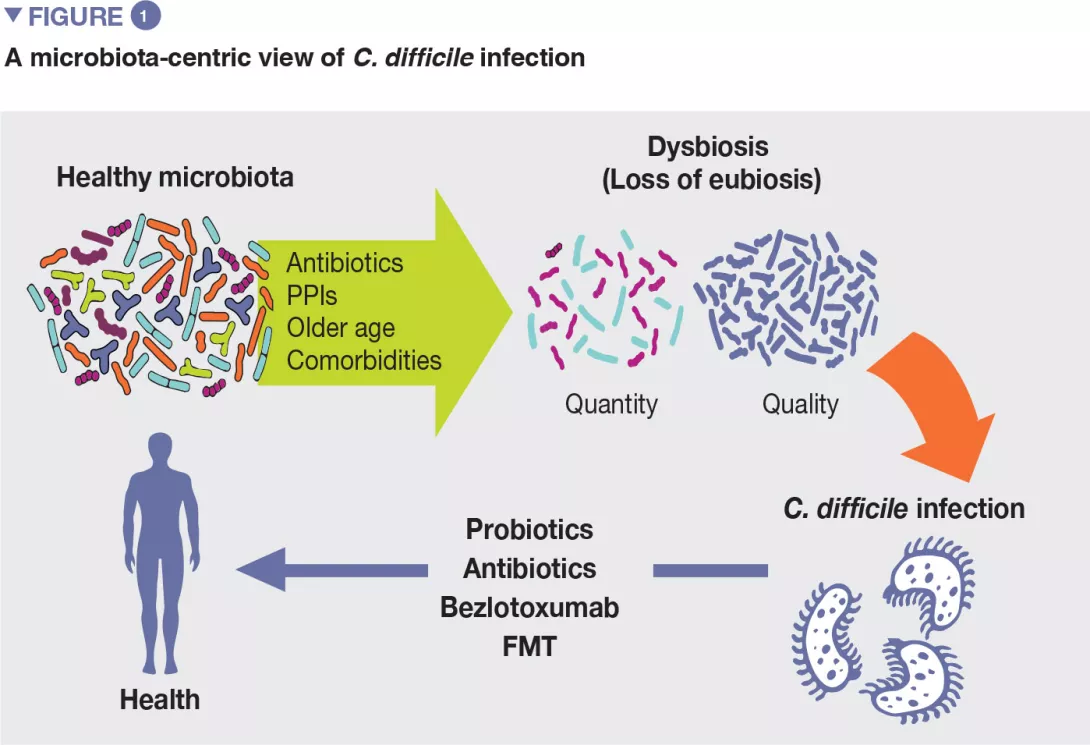
C. difficile infection (CDI) has become in recent years a clinical and socio economical burden worldwide, due to its increase in morbidity, severity, mortality, and likelihood to recur. There is a considerable involvement of gut microbiota in CDI, for many reasons. First, most risk factors associated with the development of CDI, including the overuse of broad-spectrum antibiotics or proton pump inhibitors, are associated with an imbalance of gut microbiota. Moreover, specific microbiota modulators are involved in the prevention (specific probiotics) or treatment (fecal microbiota transplantation) of CDI. In this paper, we will review epidemiology, risk factors, and approved therapies of CDI, with a microbiota-centric view.
Clostridioides difficile (C. difficile, previously called Clostridium difficile) is a gram-positive, spore-forming, obligate anaerobe. Spores allow C. difficile to persist in environments, and to be spread from infected subjects. Under specific circumstances (e.g., antibiotic-driven dysbiosis), spores are driven to germination in the large bowel, and present in a vegetative form that leads to clinical infection (Clostridium difficile infection [CDI]). In the infection phase, C. difficile produces two toxins, enterotoxin A and cytotoxin B that both cause damage to colonocytes and trigger the inflammatory response, leading to a variety of clinical pictures, from mild colitis to pseudomembranous colitis and toxic megacolon. [1]
In recent years, CDI has become a considerable healthcare and economical burden in most countries. Studies from the United States report an incidence of nearly 453,000 cases and of nearly 29,000 CDIrelated deaths in 2011, while the incidence in Europe is 124,000 cases/year, with nearly 3,700 deaths/year. Increased morbidity, hospitalization length and mortality, contribute to the considerable economic burden of CDI, which accounted for nearly $ 5 billions in the US in 2011, and for nearly € 3.7 billions in Europe in 2013. [2, 3] These figures show that the CDI incidence has risen worldwide, for several reasons. First, the increased use of antibiotics, which are a known as risk factors for CDI development. Furthermore, the spreading of specific ribotypes (mainly the virulent ribotype 027, but also the 017 in Asia, the 018 in Italy, the 17,621 in Eastern European countries, 24,422 in Oceania) has let CDI clusters develop. Additionally, there was also an increased number of diagnoses, due to the development of highly sensitive diagnostic tests (e.g., PCR), and the risen awareness of CDI among healthcare professionals. Overall, the main cause of the overall increase in CDI incidence appears to be the increased rate of recurrences. From 2001 to 2012, the annual incidence of recurrent CDI has increased by nearly 189%, while the increase in overall CDI incidence in the same time period was nearly 43%. [2] As recurrent infection is less likely than first episode to be cured by antibiotics, it is associated with longer hospitalization, increased morbidity and mortality too.
CDI is widely known to be the main cause of healthcare associated infectious diarrhoea, but recent evidence suggests that its diffusion in the community settings is growing. To date, nearly 25%-35% of CDI cases are acquired in community, probably due to several fecal-oral transmission pathways (e.g., zoonosis and food).
Despite this increase in diagnoses, the misdiagnosis/underdiagnosis of CDI is still relevant, as observed in the EUCLID study.
This finding suggests that a considerable number of patients with CDI is still not diagnosed, increasing the risk of disease diffusion.
Nosocomial CDI, a community-acquired CDI, appear to differ for several characteristics. First, nosocomial patients are more likely to present with a severe clinical picture, while community patients can even be asymptomatic carriers, increasing the risk of CDI spreading. Moreover, community- based CDI is known to spread also among patients without standard risk factors.
Although the exact pathogenic pathways of CDI are not yet clarified, several risk factors have been identified over time. [4] Their knowledge is relevant as the management of modifiable risk factors is a prevention measure against CDI. Most relevant risk factors include older age, use of antibiotics, proton pump inhibitors, and others (Figure 1).
If antibiotics remain today essential molecules in the therapeutic arsenal, it is also necessary to take into account their undesirable effects on the gut microbiota, as a considerable body of evidence supports the association between their use and many dysbiosis-associated diseases, including CDI.[5]
First, antibiotics may kill commensal bacteria that may have a direct action against C. difficile (by secreting a number of bacteriocins) and also compete with the pathogen for nutrients (e.g., sialic acid and succinate). Moreover, there is also an indirect protective role of commensal bacteria through the regulation of bile acids.
Recently, Clostridium scindens was associated with resistance to C. difficile colonisation. It has a bile acid inducible operon which is able to encode dehydroxylating enzymes that convert primary bile acids into secondary bile acids. Primary bile acids promote the germination of C. difficile spores, while secondary bile acids are able to inhibit this process.[6]
The use of systemic antibiotics is the most relevant modifiable risk factor for the development of CDI. Healthy gut microbiota can determine the successful colonisation of the large bowel by C. difficile or not, by direct and indirect pathways. In principle, the imbalance of healthy gut microbiota by broad-spectrum antibiotics may bring several consequences that drive to CDI.
As a corollary of this evidence, patients with recurrent CDI are known to have an imbalanced microbial profile, with higher relative abundance of detrimental bacterial families as Enterobacteriaceae and Veillonellaceae and lower relative abundance of beneficial families, including Ruminococcaceae, Bacteroidaceae and Lachnospiraceae.
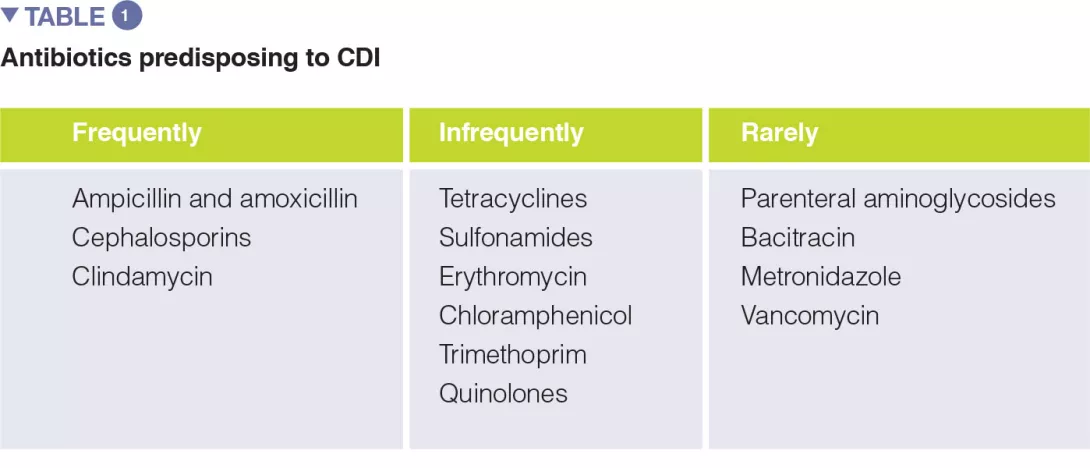
A number of systematic reviews, alone or with meta-analysis, have assessed the relevance of different antibiotic classes in CDI development. In the earliest meta-analysis (1998), antibiotics use was associated with a 6-fold increase in the risk of developing CDI, and the highest risk was observed for fluoroquinolones, clindamycin, cephalosporins. Moreover, the use of antibiotics was found to be an independent predictor of CDI recurrence (relative risk 1.76). One of the key factors to prevent CDI is represented by the antibiotic stewardship approach, so the knowledge of the CDI risk for different antibiotic classes is of paramount importance (Table 1).
The use of the following antibiotics is associated with a 2-fold higher risk of CDI among inpatients: clindamycin, cephalosporins, carbapenems, fluoroquinolones, trimethoprim, sulphonamides. In the community setting, respectively, antibiotics were found to have different risk levels for CDI development or recurrence, including: clindamycin (risk increased of 8 to 20 times), cephalosporins and fluoroquinolones (3-5 times increase), macrolides (2-3 times increase).[5]
Proton pump inhibitors (PPIs) are largely used worldwide for several upper gastrointestinal disorders, including gastroesophageal reflux disease, hiatal ernia, gastritis, H. pylori infection (together with antibiotic eradication therapy), peptic ulcer disease.
Overall PPIs are considered safe drugs. However, a large body of evidence shows that the use of PPIs is significantly associated with the development of CDI.
In principle, PPIs can increase the risk of C. difficile colonization by several pathways, including reduced acid production that can lead to small intestinal bacterial overgrowth and dysbiosis, and increase of bile salts that can promote the germination of C. difficile spores. Finally, there is no clear evidence if increased gastric pH is a safer environment for spores. [6]
The clinical evidence of a significant association between PPIs and CDI comes from several systematic reviews and meta-analyses, with odds ratios ranging from 1.26 to 2.34, based on different reports (from 3 to 67, according to different meta-analyses).
Most evidence is heterogeneous and comes from observational cohorts, so potentially confounding factors, including other drugs and co-morbidities, could reduce the quality of this finding. However, the association between PPIs and CDI kept significant even after stratification for antibiotic use, both in cohort studies and in case-control reports.
The detrimental role of PPI was found to be stronger toward community-associated CDI, suggesting that there is a chronic overuse in communities rather then in hospitals.
Specifically, PPIs have been associated not only with CDI overall, but also with recurrent CDI by several meta-analyses (including from 3 to 16 studies), with odd ratios ranging from 1.52 to 2.51, although definitions of recurrence varied significantly among studies.
Advanced age is one of the best known risk factors for primary CDI and recurrent CDI.
Established evidence shows that CDI rates are much higher in adults over 65 years than in younger population. In a meta-analysis of 33 studies, age older than 65 years was identified as an independent predictor of recurrent CDI (relative risk 1.63).
However, age is a considerable confounder, as the use of several drugs that promote CDI, such as antibiotics or PPIs, is more common in older age. Increasing evidence suggests that the microbiota of elderly patients is less healthy (in terms of reduced microbial diversity and increase of opportunistic species) than normal, supporting again the role for microbiota imbalance in CDI.[7]
The association between CDI and selected comorbidities has also been explored systematically. In a systematic review, significantly higher risk of CDI was found for inflammatory bowel disease (OR 3.72), kidney insufficiency (OR 2.64), hematologic malignancies (OR 1.75), and diabetes mellitus (OR 1.15). This was especially true for community-acquired CDI.[7]
Traditionally, metronidazole and vancomycin have been the most common treatment options for CDI, being used both as first line options, while only vancomycin was recommended, as tapered or pulsed regimen, to treat recurrent disease.[8]
However, in recent years CDI has become more cumbersome to treat. In particular metronidazole was shown to achieve lower cure rates than vancomycin, so that vancomycin has been preferred to metronidazole also in primary infection. Overall, also vancomycin is losing its efficacy, and the rates of recurrent disease have grown. Morover, hypervirulent strains of C. difficile have emerged, specifically the ribotype 027, which is less responsive to standard antibiotic therapy and is associated with more severe clinical pictures [8]
In recent years fidaxomicin, a narrow spectrum antibiotic, was shown to be superior than vancomycin in treating CDI recurrences. However, its high costs and the recent evidence of its inferiority compared with fecal microbiota transplantation (FMT) in treating recurrent CDI are potential limitations to its widespread use.[9]
Antibiotics are an extraordinary scientific discovery that saves millions of lives but their excessive and inappropriate use has now raised serious concerns for health, notably with antibiotic resistance and microbiota dysbiosis. Let’s take a look at this dedicated page:
Each year, since 2015, the WHO organizes the World AMR Awareness Week (WAAW), which aims to increase awareness of global antimicrobial resistance.
Held on 18-24 November, this campaign encourages the general public, healthcare professionals and decision-makers to use antimicrobials carefully, to prevent the further emergence of antimicrobial resistance.
Generally, probiotics are considered a reliable option to restore healthy gut microbiota after a dysbiotic event, e.g., antibiotic treatments. Overall, some probiotics are known to be effective against antibiotic-associated diarrhea (AAD), which is a common adverse event of antibiotic regimens.[10-12] In a metanalysis of 21 randomized trials, Saccharomyces boulardii decreased significantly the risk of AAD (risk ratio: 0.47).[11]
As CDI is basically a subgroup of AAD, the efficacy of probiotics in preventing CDI was then investigated. Recently, a Cochrane review has shown, in a meta-analysis of 23 trials, that probiotics are both safe and effective for preventing CDI.[13] However, only specific probiotics, including Saccharomyces boulardii, Lactobacillus casei, a mixture of L. acidophilus and Bifidobacterium bifidum, and a mixture of L. acidophilus, L. casei and L. rhamnosus, have been found to be effective in preventing primary CDI after antibiotic therapies. In particular, S. boulardii was effective in preventing CDI in a cohort of elderly hospitalized patients, with likely saving of money. Indeed, a Canadian study showed that the use of preventative probiotics was able to save $ 518/patient than usual care, and to reduce the risk of CDI.[11] However, further, larger studies are needed to confirm the role of specific probiotics in CDI prevention.
Based on this outstanding evidence, scientific societies have included FMT among the treatment options for recurrent CDI.[14, 15] FMT is also known to increase overall survival and decrease hospitalization length in patients with recurrent CDI.[16]
Although FMT has been increasingly standardized over years, is still underdiffused worldwide. Future microbiota-based approaches that will guarantee a widespread diffusion of FMT include capsulized FMT and microbiota-based drugs.
FMT is the infusion of stools from healthy donors in the gut of a recipient to cure a dysbiosis related disorders. To date, several systematic review and meta-analyses have shown that FMT is highly effetive in curing recurrent CDI (up to 90% cure rates).
CDI is a burdensome disease that occurs mainly in patients with several risk factors, most of which are associated with gut microbiota imbalance, including antibiotic overuse, proton pump inhibitors, and older age. Also from a microbiological point of view, the microbial profile of patients with CDI is characterized by a deep imbalance of gut microbiota. Therapeutic microbiota modulators have been shown to be effective in preventing (specific probiotics, some Lactobacillus strains and S. boulardii) or curing (FMT) recurrent CDI, paving the way for a microbiota-based approach for the management of this disorder.
 Antibiotic resistance is a web of several problems
Antibiotic resistance is a web of several problems
 Microbiota at the forefront of antibiotic resistance
Microbiota at the forefront of antibiotic resistance
By Prof. Markku Voutilainen
Turku University Faculty of Medicine; Turku University Hospital, Department of Gastroenterology, Turku, Finland
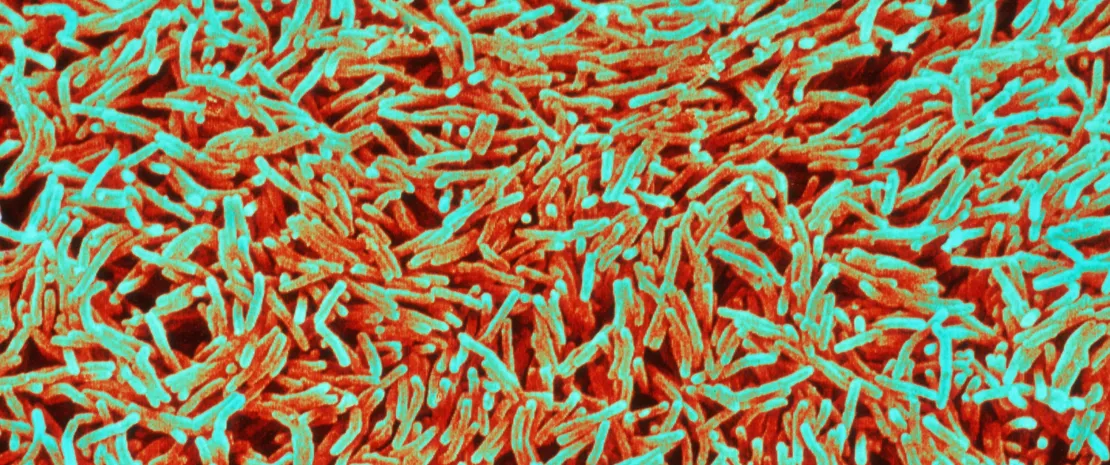
5-10% of persons infected worldwide with Mycobacterium tuberculosis (MT) will progress to active TB. Recent research highlighted that gut dysbiosis induced by treatment could be involved in the disease development by compromising immune protection against MT. This review summarizes how the gut microbiota, lung immunity could be linked during the disease; and how the gut microbiota dysbiosis induced by the protracted anti-TB antibiotics treatment is involved to an increased susceptibility to MT re-infection or TB recrudescence after successful treatment cure. The authors also indicate that the gut microbiota biosignature might help recognizing healthy from active TB patients.
Very low-carbohydrate, high-fat ketogenic diet (KD) is used in refractory pediatric epilepsy, and some evidence supports KD use in diabetes and obesity but their metabolic and immune consequences remain unclear. The authors examined the impact of KD on human and mice gut microbiota via metagenomics and metabolomics and compared with high-fat diets impact: several bifidobacterial species were reduced, and an increase of Firmicutes/Bacteroidetes ratio induced by high-fat diet reversed. Increased plasma β-hydroxybutyrate levels inhibit bifidobacterial growth. KD reduced proinflammatory Th17 cell accumulation in mice adipose tissue and inhibited induction of intestinal Th17 cells.
By Prof. Markku Voutilainen
Turku University Faculty of Medicine; Turku University Hospital, Department of Gastroenterology, Turku, Finland
Kim HS, Yosipovitch G. The Skin Microbiota and Itch: Is There a Link? J Clin Med 2020 ; 9 : 1190.
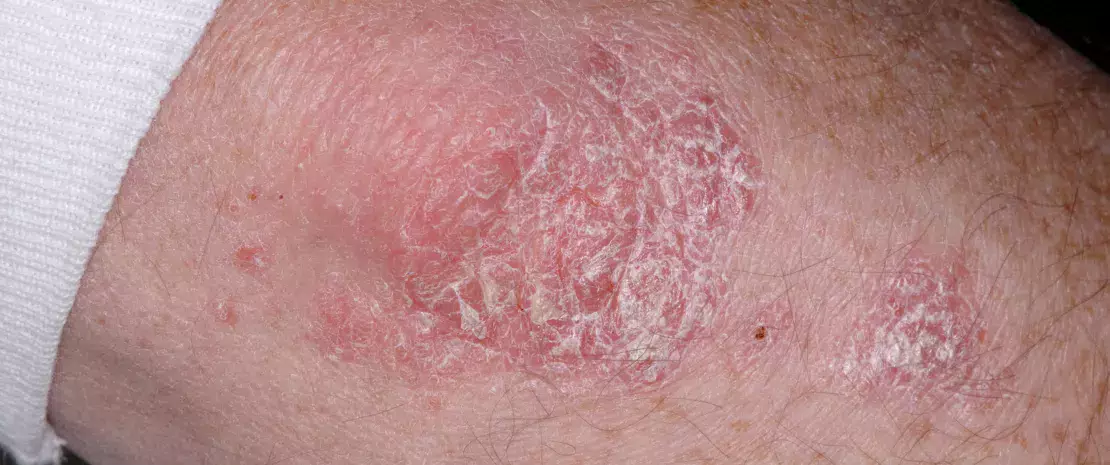
The authors discuss the role of skin microbiota in the pathogenesis of itch. Itch sensation is mediated via epidermal nerve fibres (pruriceptors) driving by chemical mediators that originate from a complex interaction between keratinocytes (KC), inflammatory cells, nerve endings and the skin microbiota, relaying itch signals to the brain. Skin dysbiosis is characterized by production of proteases, pathogen-associated molecular patterns, and toxins, leading to skin barrier damage. Mast cell degranulation induced by delta-toxin prompt inflammation and itching. Skin microbiota and brain communicate via neurochemicals (acetylcholine, histamine, catecholamines, corticotropin) originate from skin microbiota. Stress intensifies itch via the skin-brain axis, where the amygdala seems to modulate itching sensation via microbial signals. Chronic stress increases cortisol production, directly activates skin bacteria by increasing the virulence of skin pathogens, leading to a weakening of the skin barrier and to an aggravation of the itch sensation. The authors conclude that cosmetics/transdermal drugs that modulate skin microbiota might have the potential to ameliorate itch.
The authors examined the microbiota in psoriatic lesions and unaffected skin in psoriasis vulgaris (PS) patients and healthy controls by quantitative PCR and 16S rRNA sequencing. Higher bacterial load and lower diversity was observed in PS lesions than patients unaffected and controls’ skin. Cutibacterium (Cu) was reduced in lesions, whereas Corynebacterium (Cr) was increased. Compared with patients’ unaffected skin, Cr/Cu + Cr ratio was higher in the lesions. These findings indicate that PS was the major cause for the imbalance between Cu and Cr between lesions and unaffected skin or controls. Cr load correlated with the severity of PS lesions, whereas Cu load showed correlation with the abnormity of skin capacitance. The present study suggests that skin microbiota might play a significant role in the pathogenesis of PS.
By Prof. Markku Voutilainen
Turku University Faculty of Medicine; Turku University Hospital, Department of Gastroenterology, Turku, Finland
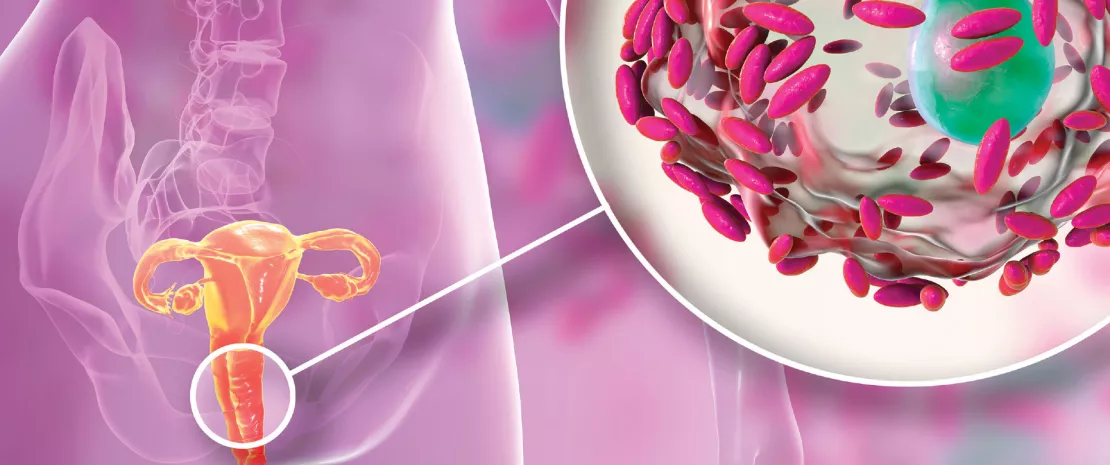
Recurrent implantation failure (RIF) is defined as a failure to achieve a clinical pregnancy after transfer of at least four good-quality embryos in a woman under the age of 40 years. Embryonal and uterine factors or maternal systemic diseases may cause RIF, but some women do not have recognizable etiology. The authors focused on the vaginal microbiota and metabolome of women with RIF. They found that RIF patients suffered vaginal dysbiosis, having a more diverse and abundant bacteria with an increase of many anaerobic and aerobic bacteria which could be linked to bacterial vaginosis and aerobic vaginitis or urinary tract infection, respectively. Conversely, at genus level their vaginal microbiota was decreased in Lactobacillus (LB); at species level L. iners was reduced and L. crispatus was the most abundant species in the RIF group. Increased vaginal bacterial diversity, LB depletion and related metabolic changes could serve as biomarkers capable of predicting the risk of RIF.
This review summarizes the role played by vaginal microbiota in urinary tract infections (UTI) as mounting evidence indicates that vagina may serve as a reservoir for uropathogens and increase susceptibility to UTI. Escherichia coli is the commonest cause of UTI and can colonize the vagina, which can be increased if the vaginal Lactobacillus (LB) colonization is reduced. Some vaginal bacteria are frequently detected in the urine but are underappreciated as uropathogens, because they are difficult to detect in routine clinical practice. For example, bacterial vaginosis (BV) is characterized by Gram-negative anaerobes, species belonging to Actinobacteria and Firmicutes phyla while LB is reduced and BV patients have a higher UTI risk. Gardnerella vaginalis is detected in BV and can cause acute or recurrent UTI. Group B Streptococcus may cause both aerobic vaginitis and UTI. Finally, some vaginal bacteria may enter the urinary tract and can transit briefly, cause immunomodulation or injury and unbalanced the host-pathogen interactions to influence the outcomes of uropathogenesis.
Congress review
By Dr Dragos Ciocan
CCA Hepato-gastroenterology and nutrition department Antoine-Béclère Hospital, Clamart, France

Despite the Covid-19 pandemic, this year we were able to benefit from an on-line version of the Journees francophones d’hépato-gastroenterologie (French-speaking hepato-gastroenterology congress) with the advantage of being able to access presentations recorded between 3 and 20 July 2020. These eJFHOD reached a total of 7,924 users, and 172,937 pages were viewed. As each year, original studies on the intestinal microbiota (IM) were presented at this congress.
Third most frequent cancer in humans, sporadic colorectal cancer (CRC) develops following interactions between the host and its environment, and the IM is thought to be implicated [1]. Professor Sobhani presented the results of a study which investigated the links between epigenetic mechanisms promoted by bacteria of the IM and the onset of CRC [2]. Mice transplanted with faecal samples from patients with CRC developed precancerous colonic lesions, associated with an increase in hypermethylated genes. Donors with CRC exhibited methylation anomalies of several gene promoters associated with intestinal dysbiosis. Using the identified microbial and epigenetic signatures, a pilot study (n = 266) was conducted in humans in order to develop a blood test for the diagnosis of CRC. A cumulative methylation index (CMI, measuring the hypermethylation level of 3 genes) was identified as a predictive factor in the onset of CRC. These results were validated in a prospective cohort of 1,000 patients. Intestinal dysbiosis in patients with a positive CMI was characterised by an increase in pro-methylating bacterial species. This work indicates that intestinal dysbiosis associated with CRC could promote colon carcinogenesis via deregulation of the methylation of certain genes. The cumulative hypermethylation index (CMI) and/or pro-methylating bacteria are thus potential biomarkers for CRC diagnosis, or be used in the evaluation of the effects of treatments modulating the IM in patients with CRC.
In a study coordinated by Professor Seksik, the authors studied the role of MAM (microbial anti-inflammatory molecule, produced by Faecalibacterium prausnitzii and reduced in patients with Crohn’s disease, CD [3]) as a biomarker of intestinal dysbiosis and diagnostic aid in CD. The authors showed that loss of MAM is associated with the diagnosis of CD. This preliminary study in a small number of patients (24 patients in relapse, 24 in remission and 12 healthy controls) paves the way to the diagnosis of CD based on the IM, but these preliminary results require validation in independent cohorts.
It is known that bacteria detect and respond to environmental signals (an ability called Quorum Sensing). Of the molecules which are part of this system, 3-oxo-C12:2 is low in patients with chronic inflammatory bowel disease (IBD), this reduction appears to be correlated with the observed intestinal dysbiosis [4]. In a study presented by D. Aguanno, the authors studied the impact of this molecule on the epithelial cells of the intestine and showed that this did not modify paracellular permeability but attenuated the deleterious effects on the tight junctions induced by pro-inflammatory cytokines. In a second study, Coquant et al. showed that 3-oxo-C12:2 exerted an anti-inflammatory effect on immunoreactive cells, partly mediated by the T2R138 receptor. This molecule may therefore have protective effects on the intestinal barrier, modulate the inflammatory response and thus represent a novel therapeutic perspective in IBD.
