Live probiotic coconut yogurt, is it worth recommending?
By Dr. Hanna Stolińska
Dietetic Clinic, Warsaw, Poland

Health influencers on social media are buzzing about a super-live probiotic coconut yogurt claiming to revolutionize gut health. Marketed as a superfood packed with billions of probiotics, it has gained a cult-like following among wellness enthusiasts. Fans praise its supposed benefits, from improved digestion to healthier skin, but what does science have to say compared to traditional probiotics? Is this coconut-derived probiotic-rich formula a true microbiome booster or just another overhyped wellness trend?
What is inside the probiotic coconut-based yogurt?
The coconut-based yogurt is made from organic coconut meat and coconut water, fermented with 16 custom probiotic strains, including Lactobacillus acidophilus, Bifidobacterium breve, and Streptococcus thermophilus.
What health benefits does it claim to offer? Is it truly beneficial for health?
It is reported online that consuming this yogurt may improve digestion, reduce bloating, promote healthier skin, and strengthen the immune system. While probiotics can offer health benefits, scientific evidence specifically supporting this yogurt’s claim remains limited.
We know that coconut flesh is mainly composed of lipids and such yogurt provides very little protein. On top of that coconut oil is used as a cure for all sorts of ailments, such as fighting viruses and bacteria, supporting immunity, reducing cholesterol, supporting thyroid function, and even weight loss 1-4.
Coconut oil contains medium-chain triglycerides (MCT) fatty acids, which are more easily digested and less absorbed compared to longer-chain fatty acids. However, of all the saturated fatty acids in coconut oil, MCTs constitute only half. Some MCTs, such as lauric acid and capric acid, have antifungal and antiviral properties 1-3, but the purpose of consuming food is to provide components that strengthen the immune system, which then fights microbes 4.
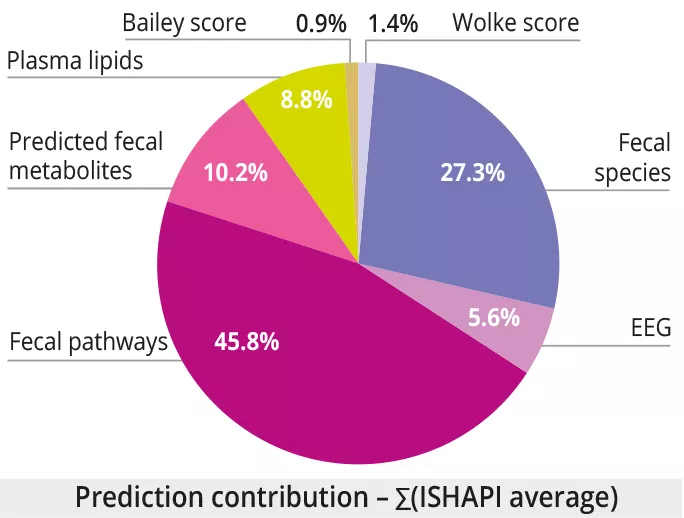
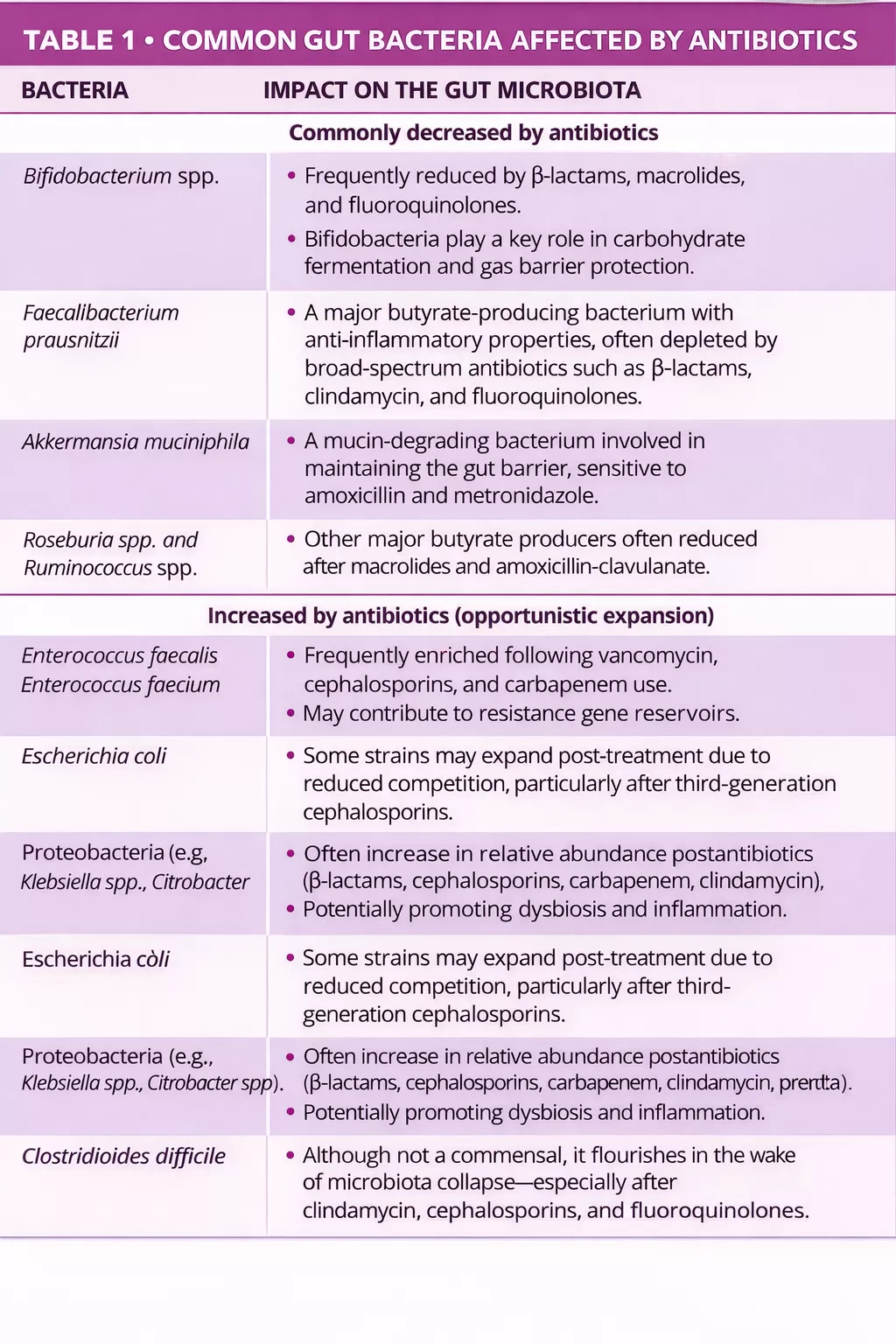
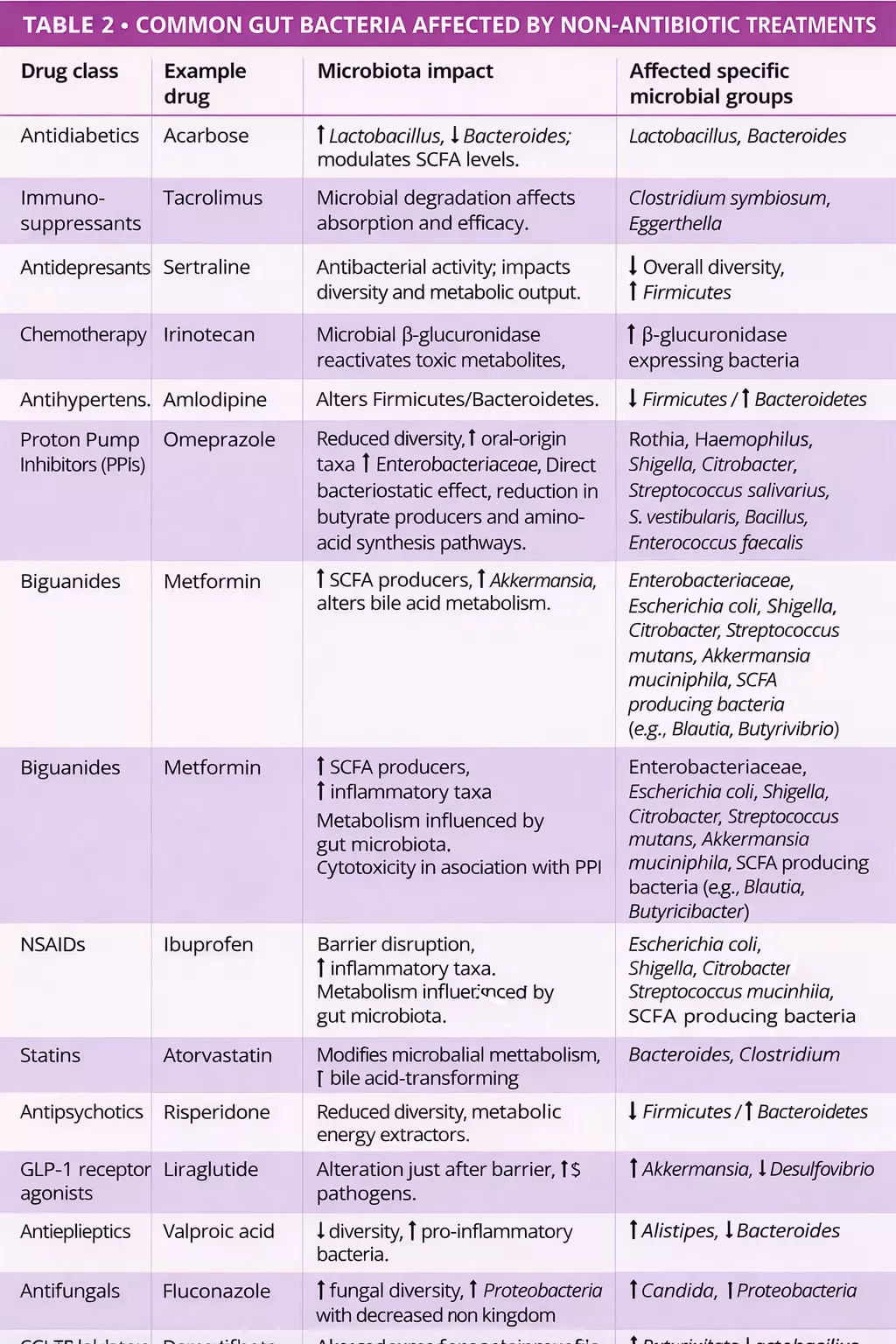
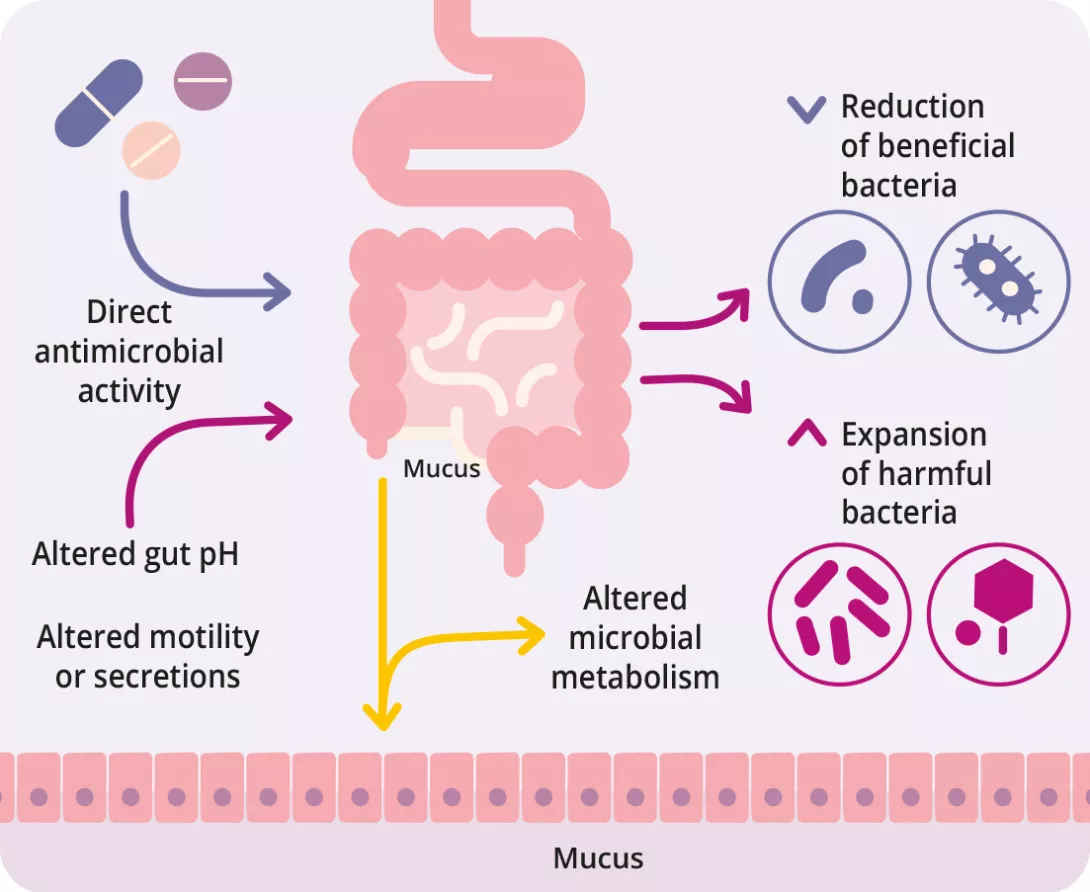
How does it impact the gut microbiota?
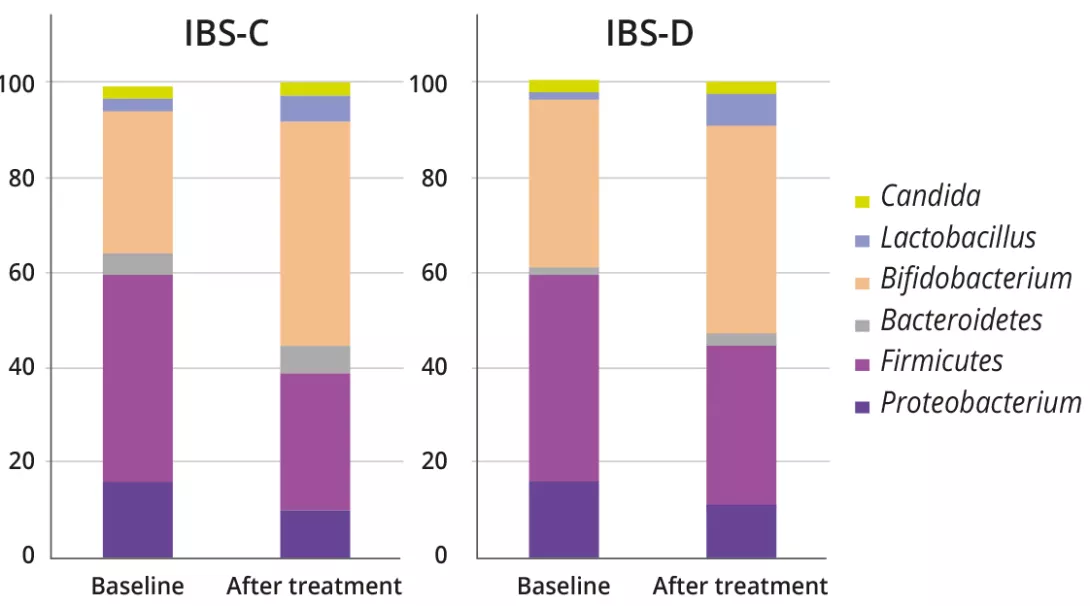
This probiotic-enriched yogurt contains live bacteria that can potentially influence gut microbiota. However, the effectiveness of probiotics depends on various factors, including the specific strains used, their ability to survive stomach acid, and the individual’s existing gut microbiome composition.
Some studies suggest that mediumchain triglycerides (MCTs), found in coconut, may affect microbiome composition. However, research on probiotic coconut-based products is still in its early stages. Moreover, while coconut oil contains antimicrobial compounds like lauric acid (converted into monolaurin in the body), this does not necessarily translate into overall gut health benefits 1, 3.
An interesting rat study investigating different dietary oils’ effects on gut microbiota found that coconut oil consumption led to reduced bacterial diversity, increased markers of metabolic endotoxemia, fatty liver disease, and higher LDL cholesterol levels 5. While animal studies provide insight, further clinical trials in humans are needed to determine this probiotic-enriched yogurt’s actual effects on gut health.
From your dietary perspective, are these new probiotic products worth considering?
Remember that food is primarily intended to provide us with nutrients, vitamins and minerals. Coconut dairy, like yogurt, does not have high nutritional density 1, 3.
While this probiotic-rich yogurt presents an innovative approach to delivering beneficial bacteria, its high-fat content, low protein levels, and cost should be considered when recommending it as a dietary option 2, 6. Traditional probiotic-rich foods, such as yogurt, kefir, and fermented vegetables, provide similar benefits with a more balanced nutritional profile. A varied and balanced diet, low in processed foods and rich in vegetables, fruits, whole grains, and legumes, supports microbiome diversity.
- Boateng L, Ansong R, Owusu WB. Coconut oil and palm oil’s role in nutrition, health and national development: A review. Ghana Med J 2016; 50: 189-96.
- Eyres L, Eyres MF, Chisholm A, Brown RC. Coconut oil consumption and cardiovascular risk factors in humans. Nutr Rev 2016; 74: 267-80.
- Lockyer S, Stanner S. Coconut oil – a nutty idea? British Nutrition Foundation Nutrition Bulletin 2016; 41: 42-54.
- Joshi S, Kaushik V, Gode V, Mhaskar S. Coconut Oil and Immunity: What do we really know about it so far? J Assoc Physicians India 2020; 68: 67-72.
- López-Salazar V, Tapia MS, Tobón-Cornejo S, Díaz D, Alemán-Escondrillas G, Granados-Portillo O, Noriega L, Tovar AR, Torres N. Consumption of soybean or olive oil at recommended concentrations increased the intestinal microbiota diversity and insulin sensitivity and prevented fatty liver compared to the effects of coconut oil. J Nutr Biochem. 2021; 94: 108751.
- Sacks FM, Lichtenstein AH, Wu JHY, et al.; American Heart Association. Dietary Fats and Cardiovascular Disease: A Presidential Advisory From the American Heart Association. Circulation 2017; 136: e1-e23.